
| Cancer | Chr | Position | Mutation Type | dbSNP | Protein-change | Allele Freq | RBD |
|---|---|---|---|---|---|---|---|
| ACC | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| ESCA | |||||||
| ESCA | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| HNSC | |||||||
| HNSC | |||||||
| LAML | |||||||
| LGG | |||||||
| LGG | |||||||
| LGG | |||||||
| LIHC | |||||||
| LIHC | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| OV | |||||||
| PAAD | |||||||
| READ | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCS | |||||||
| UCS |
| Cancer | Type | Freq | Q-value |
|---|---|---|---|
| KIRC | |||
| THCA |
| Cancer | P-value | Q-value |
|---|---|---|
| KIRC | ||
| STAD | ||
| SARC | ||
| MESO | ||
| HNSC | ||
| BRCA | ||
| ESCA | ||
| TGCT | ||
| PAAD | ||
| LAML | ||
| KICH | ||
| LIHC | ||
| OV |
| Protein ID | Domain | Pfam ID | E-value | Domain number | Total number |
|---|---|---|---|---|---|
| ENSP00000362063 | RRM_1 | PF00076.22 | 6e-25 | 1 | 1 |
| ENSP00000387996 | RRM_1 | PF00076.22 | 6.2e-25 | 1 | 1 |
| ENSP00000415705 | RRM_1 | PF00076.22 | 2.4e-24 | 1 | 1 |
| Transcript ID | Name | Length | RefSeq ID | Protein ID | Length | RefSeq ID | UniportKB ID |
|---|---|---|---|---|---|---|---|
| ENST00000475126 | CSTF2-204 | 1693 | - | ENSP00000432060 | 438 (aa) | - | E9PID8 |
| ENST00000413437 | CSTF2-202 | 757 | - | ENSP00000415705 | 225 (aa) | - | A0A0A0MT56 |
| ENST00000415585 | CSTF2-203 | 2074 | - | ENSP00000387996 | 597 (aa) | - | E7EWR4 |
| ENST00000372972 | CSTF2-201 | 1958 | - | ENSP00000362063 | 577 (aa) | - | P33240 |
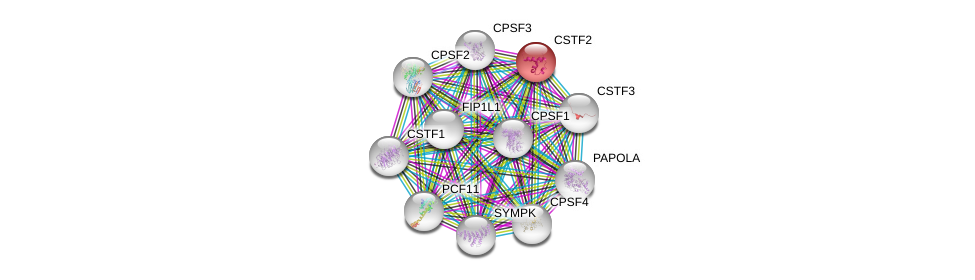
| Pathway ID | Pathway Name | Source |
|---|---|---|
| hsa03015 | mRNA surveillance pathway | KEGG |
| Ensembl ID | Gene Symbol | Coverage | Identiy | Paralog | Gene Symbol | Coverage | Identiy |
|---|---|---|---|---|---|---|---|
| ENSG00000101811 | CSTF2 | 84 | 88.360 | ENSG00000177613 | CSTF2T | 99 | 69.255 |
| ENSG00000101811 | CSTF2 | 57 | 33.333 | ENSG00000179950 | PUF60 | 63 | 33.333 |
| Ensembl ID | Gene Symbol | Coverage | Identiy | Ortholog | Gene Symbol | Coverage | Identiy | Species |
|---|---|---|---|---|---|---|---|---|
| ENSG00000101811 | CSTF2 | 95 | 83.178 | ENSAPOG00000005630 | cstf2 | 97 | 60.561 | Acanthochromis_polyacanthus |
| ENSG00000101811 | CSTF2 | 100 | 95.861 | ENSAMEG00000002082 | CSTF2 | 100 | 95.861 | Ailuropoda_melanoleuca |
| ENSG00000101811 | CSTF2 | 81 | 92.350 | ENSACIG00000012852 | cstf2 | 96 | 58.160 | Amphilophus_citrinellus |
| ENSG00000101811 | CSTF2 | 81 | 92.350 | ENSAOCG00000020905 | cstf2 | 97 | 60.513 | Amphiprion_ocellaris |
| ENSG00000101811 | CSTF2 | 81 | 92.350 | ENSAPEG00000006526 | cstf2 | 97 | 60.513 | Amphiprion_percula |
| ENSG00000101811 | CSTF2 | 79 | 93.820 | ENSATEG00000018186 | cstf2 | 97 | 57.541 | Anabas_testudineus |
| ENSG00000101811 | CSTF2 | 84 | 94.180 | ENSAPLG00000007255 | CSTF2 | 98 | 78.487 | Anas_platyrhynchos |
| ENSG00000101811 | CSTF2 | 100 | 98.614 | ENSANAG00000022342 | CSTF2 | 100 | 98.614 | Aotus_nancymaae |
| ENSG00000101811 | CSTF2 | 81 | 93.407 | ENSACLG00000000779 | cstf2 | 93 | 60.377 | Astatotilapia_calliptera |
| ENSG00000101811 | CSTF2 | 94 | 82.464 | ENSAMXG00000021477 | cstf2 | 96 | 98.413 | Astyanax_mexicanus |
| ENSG00000101811 | CSTF2 | 100 | 95.147 | ENSBTAG00000003547 | CSTF2 | 100 | 95.147 | Bos_taurus |
| ENSG00000101811 | CSTF2 | 100 | 98.995 | ENSCJAG00000004422 | CSTF2 | 100 | 98.995 | Callithrix_jacchus |
| ENSG00000101811 | CSTF2 | 100 | 96.360 | ENSCAFG00000017551 | CSTF2 | 100 | 96.360 | Canis_familiaris |
| ENSG00000101811 | CSTF2 | 100 | 96.482 | ENSCAFG00020007619 | CSTF2 | 100 | 96.482 | Canis_lupus_dingo |
| ENSG00000101811 | CSTF2 | 100 | 95.494 | ENSCHIG00000016994 | CSTF2 | 100 | 95.494 | Capra_hircus |
| ENSG00000101811 | CSTF2 | 100 | 98.325 | ENSTSYG00000006529 | CSTF2 | 100 | 98.325 | Carlito_syrichta |
| ENSG00000101811 | CSTF2 | 100 | 96.147 | ENSCPOG00000015255 | CSTF2 | 100 | 96.147 | Cavia_porcellus |
| ENSG00000101811 | CSTF2 | 100 | 98.995 | ENSCCAG00000032975 | CSTF2 | 100 | 98.995 | Cebus_capucinus |
| ENSG00000101811 | CSTF2 | 100 | 99.497 | ENSCATG00000022835 | CSTF2 | 100 | 99.497 | Cercocebus_atys |
| ENSG00000101811 | CSTF2 | 100 | 95.980 | ENSCLAG00000001904 | CSTF2 | 100 | 95.980 | Chinchilla_lanigera |
| ENSG00000101811 | CSTF2 | 100 | 99.665 | ENSCSAG00000010360 | CSTF2 | 100 | 99.665 | Chlorocebus_sabaeus |
| ENSG00000101811 | CSTF2 | 71 | 100.000 | ENSCHOG00000012803 | CSTF2 | 75 | 100.000 | Choloepus_hoffmanni |
| ENSG00000101811 | CSTF2 | 96 | 87.558 | ENSCPBG00000014293 | CSTF2 | 100 | 76.425 | Chrysemys_picta_bellii |
| ENSG00000101811 | CSTF2 | 100 | 99.665 | ENSCANG00000030612 | CSTF2 | 100 | 99.665 | Colobus_angolensis_palliatus |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | ENSCGRG00001023023 | Cstf2 | 100 | 92.931 | Cricetulus_griseus_chok1gshd |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | ENSCGRG00000003897 | Cstf2 | 100 | 89.828 | Cricetulus_griseus_crigri |
| ENSG00000101811 | CSTF2 | 83 | 91.398 | ENSCVAG00000005731 | cstf2 | 96 | 60.732 | Cyprinodon_variegatus |
| ENSG00000101811 | CSTF2 | 86 | 88.660 | ENSDARG00000090788 | cstf2 | 93 | 92.188 | Danio_rerio |
| ENSG00000101811 | CSTF2 | 100 | 92.746 | ENSDORG00000007195 | Cstf2 | 100 | 92.746 | Dipodomys_ordii |
| ENSG00000101811 | CSTF2 | 96 | 96.279 | ENSETEG00000004931 | CSTF2 | 68 | 93.243 | Echinops_telfairi |
| ENSG00000101811 | CSTF2 | 100 | 96.985 | ENSEASG00005001216 | CSTF2 | 100 | 96.985 | Equus_asinus_asinus |
| ENSG00000101811 | CSTF2 | 100 | 96.985 | ENSECAG00000009325 | CSTF2 | 100 | 96.985 | Equus_caballus |
| ENSG00000101811 | CSTF2 | 100 | 96.817 | ENSFCAG00000007000 | CSTF2 | 100 | 96.817 | Felis_catus |
| ENSG00000101811 | CSTF2 | 92 | 77.837 | ENSFALG00000003859 | CSTF2 | 100 | 78.014 | Ficedula_albicollis |
| ENSG00000101811 | CSTF2 | 100 | 96.360 | ENSFDAG00000002416 | CSTF2 | 100 | 96.360 | Fukomys_damarensis |
| ENSG00000101811 | CSTF2 | 82 | 90.811 | ENSFHEG00000017065 | cstf2 | 93 | 61.207 | Fundulus_heteroclitus |
| ENSG00000101811 | CSTF2 | 81 | 88.462 | ENSGMOG00000005145 | cstf2 | 99 | 56.799 | Gadus_morhua |
| ENSG00000101811 | CSTF2 | 100 | 79.181 | ENSGALG00000006760 | CSTF2 | 100 | 85.667 | Gallus_gallus |
| ENSG00000101811 | CSTF2 | 81 | 91.257 | ENSGAFG00000020027 | cstf2 | 96 | 59.520 | Gambusia_affinis |
| ENSG00000101811 | CSTF2 | 85 | 86.667 | ENSGACG00000017326 | cstf2 | 93 | 81.633 | Gasterosteus_aculeatus |
| ENSG00000101811 | CSTF2 | 100 | 78.628 | ENSGAGG00000006037 | CSTF2 | 100 | 78.515 | Gopherus_agassizii |
| ENSG00000101811 | CSTF2 | 100 | 100.000 | ENSGGOG00000022267 | CSTF2 | 100 | 100.000 | Gorilla_gorilla |
| ENSG00000101811 | CSTF2 | 81 | 93.407 | ENSHBUG00000016183 | cstf2 | 93 | 60.309 | Haplochromis_burtoni |
| ENSG00000101811 | CSTF2 | 100 | 82.322 | ENSHGLG00000013455 | - | 100 | 82.322 | Heterocephalus_glaber_female |
| ENSG00000101811 | CSTF2 | 100 | 97.054 | ENSHGLG00000000659 | - | 100 | 97.054 | Heterocephalus_glaber_female |
| ENSG00000101811 | CSTF2 | 100 | 82.496 | ENSHGLG00100015391 | - | 100 | 82.496 | Heterocephalus_glaber_male |
| ENSG00000101811 | CSTF2 | 100 | 97.054 | ENSHGLG00100001130 | - | 100 | 97.054 | Heterocephalus_glaber_male |
| ENSG00000101811 | CSTF2 | 93 | 82.629 | ENSHCOG00000005157 | cstf2 | 97 | 60.554 | Hippocampus_comes |
| ENSG00000101811 | CSTF2 | 80 | 91.713 | ENSIPUG00000017839 | cstf2 | 99 | 60.956 | Ictalurus_punctatus |
| ENSG00000101811 | CSTF2 | 100 | 96.187 | ENSSTOG00000023445 | CSTF2 | 100 | 96.187 | Ictidomys_tridecemlineatus |
| ENSG00000101811 | CSTF2 | 100 | 94.801 | ENSJJAG00000020507 | Cstf2 | 100 | 94.801 | Jaculus_jaculus |
| ENSG00000101811 | CSTF2 | 81 | 91.758 | ENSKMAG00000001068 | cstf2 | 96 | 59.722 | Kryptolebias_marmoratus |
| ENSG00000101811 | CSTF2 | 54 | 94.915 | ENSLBEG00000028720 | cstf2 | 87 | 89.062 | Labrus_bergylta |
| ENSG00000101811 | CSTF2 | 93 | 84.360 | ENSLACG00000015927 | CSTF2 | 99 | 67.805 | Latimeria_chalumnae |
| ENSG00000101811 | CSTF2 | 98 | 77.778 | ENSLOCG00000015300 | cstf2 | 93 | 64.505 | Lepisosteus_oculatus |
| ENSG00000101811 | CSTF2 | 96 | 97.674 | ENSLAFG00000000474 | CSTF2 | 100 | 92.295 | Loxodonta_africana |
| ENSG00000101811 | CSTF2 | 100 | 99.665 | ENSMFAG00000043888 | CSTF2 | 100 | 99.665 | Macaca_fascicularis |
| ENSG00000101811 | CSTF2 | 100 | 98.827 | ENSMMUG00000013008 | CSTF2 | 100 | 98.827 | Macaca_mulatta |
| ENSG00000101811 | CSTF2 | 100 | 99.665 | ENSMNEG00000041824 | CSTF2 | 100 | 99.665 | Macaca_nemestrina |
| ENSG00000101811 | CSTF2 | 100 | 99.665 | ENSMLEG00000036870 | CSTF2 | 100 | 99.665 | Mandrillus_leucophaeus |
| ENSG00000101811 | CSTF2 | 81 | 92.308 | ENSMAMG00000020364 | cstf2 | 96 | 58.834 | Mastacembelus_armatus |
| ENSG00000101811 | CSTF2 | 81 | 93.407 | ENSMZEG00005007200 | cstf2 | 96 | 60.377 | Maylandia_zebra |
| ENSG00000101811 | CSTF2 | 84 | 86.243 | ENSMGAG00000004139 | CSTF2 | 100 | 85.259 | Meleagris_gallopavo |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | ENSMAUG00000000965 | Cstf2 | 100 | 93.276 | Mesocricetus_auratus |
| ENSG00000101811 | CSTF2 | 100 | 96.817 | ENSMICG00000014121 | CSTF2 | 100 | 96.817 | Microcebus_murinus |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | ENSMOCG00000008081 | Cstf2 | 100 | 91.897 | Microtus_ochrogaster |
| ENSG00000101811 | CSTF2 | 84 | 92.308 | ENSMMOG00000000538 | cstf2 | 97 | 54.096 | Mola_mola |
| ENSG00000101811 | CSTF2 | 96 | 88.372 | ENSMODG00000011785 | CSTF2 | 100 | 82.178 | Monodelphis_domestica |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | MGP_CAROLIEiJ_G0033383 | Cstf2 | 100 | 93.276 | Mus_caroli |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | ENSMUSG00000031256 | Cstf2 | 100 | 93.276 | Mus_musculus |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | MGP_PahariEiJ_G0031923 | Cstf2 | 100 | 93.437 | Mus_pahari |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | MGP_SPRETEiJ_G0034548 | Cstf2 | 100 | 93.276 | Mus_spretus |
| ENSG00000101811 | CSTF2 | 96 | 98.605 | ENSMPUG00000002339 | CSTF2 | 100 | 91.111 | Mustela_putorius_furo |
| ENSG00000101811 | CSTF2 | 96 | 95.814 | ENSMLUG00000004598 | CSTF2 | 98 | 89.348 | Myotis_lucifugus |
| ENSG00000101811 | CSTF2 | 100 | 95.164 | ENSNGAG00000021985 | Cstf2 | 100 | 95.337 | Nannospalax_galili |
| ENSG00000101811 | CSTF2 | 80 | 92.265 | ENSNBRG00000021694 | cstf2 | 100 | 56.306 | Neolamprologus_brichardi |
| ENSG00000101811 | CSTF2 | 100 | 99.665 | ENSNLEG00000007510 | CSTF2 | 100 | 99.665 | Nomascus_leucogenys |
| ENSG00000101811 | CSTF2 | 91 | 79.848 | ENSMEUG00000014025 | - | 100 | 81.369 | Notamacropus_eugenii |
| ENSG00000101811 | CSTF2 | 96 | 87.442 | ENSMEUG00000007875 | - | 100 | 79.376 | Notamacropus_eugenii |
| ENSG00000101811 | CSTF2 | 96 | 88.837 | ENSMEUG00000011186 | - | 100 | 84.055 | Notamacropus_eugenii |
| ENSG00000101811 | CSTF2 | 78 | 96.571 | ENSOPRG00000003106 | CSTF2 | 87 | 80.995 | Ochotona_princeps |
| ENSG00000101811 | CSTF2 | 100 | 95.494 | ENSODEG00000002059 | CSTF2 | 100 | 95.494 | Octodon_degus |
| ENSG00000101811 | CSTF2 | 81 | 92.857 | ENSONIG00000002631 | cstf2 | 99 | 57.477 | Oreochromis_niloticus |
| ENSG00000101811 | CSTF2 | 96 | 89.767 | ENSOANG00000004208 | CSTF2 | 87 | 90.000 | Ornithorhynchus_anatinus |
| ENSG00000101811 | CSTF2 | 100 | 95.321 | ENSOCUG00000010468 | CSTF2 | 100 | 95.321 | Oryctolagus_cuniculus |
| ENSG00000101811 | CSTF2 | 83 | 90.860 | ENSORLG00000001380 | cstf2 | 97 | 58.120 | Oryzias_latipes |
| ENSG00000101811 | CSTF2 | 83 | 90.860 | ENSORLG00020021917 | cstf2 | 97 | 58.462 | Oryzias_latipes_hni |
| ENSG00000101811 | CSTF2 | 83 | 90.860 | ENSORLG00015011996 | cstf2 | 97 | 58.120 | Oryzias_latipes_hsok |
| ENSG00000101811 | CSTF2 | 81 | 92.350 | ENSOMEG00000015507 | cstf2 | 97 | 59.966 | Oryzias_melastigma |
| ENSG00000101811 | CSTF2 | 96 | 98.605 | ENSOGAG00000011192 | CSTF2 | 100 | 90.143 | Otolemur_garnettii |
| ENSG00000101811 | CSTF2 | 100 | 95.494 | ENSOARG00000001022 | CSTF2 | 100 | 95.494 | Ovis_aries |
| ENSG00000101811 | CSTF2 | 100 | 100.000 | ENSPPAG00000031804 | CSTF2 | 100 | 100.000 | Pan_paniscus |
| ENSG00000101811 | CSTF2 | 100 | 96.985 | ENSPPRG00000022492 | CSTF2 | 100 | 96.985 | Panthera_pardus |
| ENSG00000101811 | CSTF2 | 100 | 96.650 | ENSPTIG00000009922 | CSTF2 | 100 | 96.650 | Panthera_tigris_altaica |
| ENSG00000101811 | CSTF2 | 100 | 100.000 | ENSPTRG00000022090 | CSTF2 | 100 | 100.000 | Pan_troglodytes |
| ENSG00000101811 | CSTF2 | 100 | 99.665 | ENSPANG00000002303 | CSTF2 | 100 | 99.665 | Papio_anubis |
| ENSG00000101811 | CSTF2 | 87 | 89.744 | ENSPKIG00000005033 | cstf2 | 97 | 61.073 | Paramormyrops_kingsleyae |
| ENSG00000101811 | CSTF2 | 81 | 92.308 | ENSPMGG00000006534 | cstf2 | 97 | 58.814 | Periophthalmus_magnuspinnatus |
| ENSG00000101811 | CSTF2 | 96 | 98.605 | ENSPEMG00000019415 | Cstf2 | 100 | 92.241 | Peromyscus_maniculatus_bairdii |
| ENSG00000101811 | CSTF2 | 96 | 88.837 | ENSPCIG00000002989 | CSTF2 | 100 | 82.759 | Phascolarctos_cinereus |
| ENSG00000101811 | CSTF2 | 81 | 91.803 | ENSPLAG00000011176 | cstf2 | 96 | 57.333 | Poecilia_latipinna |
| ENSG00000101811 | CSTF2 | 81 | 91.803 | ENSPMEG00000017743 | cstf2 | 96 | 57.167 | Poecilia_mexicana |
| ENSG00000101811 | CSTF2 | 81 | 91.803 | ENSPREG00000011481 | cstf2 | 97 | 60.272 | Poecilia_reticulata |
| ENSG00000101811 | CSTF2 | 100 | 99.832 | ENSPPYG00000020541 | CSTF2 | 100 | 99.832 | Pongo_abelii |
| ENSG00000101811 | CSTF2 | 100 | 97.822 | ENSPCOG00000027744 | CSTF2 | 100 | 97.822 | Propithecus_coquereli |
| ENSG00000101811 | CSTF2 | 97 | 80.790 | ENSPVAG00000008342 | CSTF2 | 100 | 80.790 | Pteropus_vampyrus |
| ENSG00000101811 | CSTF2 | 81 | 93.407 | ENSPNYG00000010852 | cstf2 | 94 | 91.781 | Pundamilia_nyererei |
| ENSG00000101811 | CSTF2 | 95 | 82.629 | ENSPNAG00000018887 | cstf2 | 97 | 61.873 | Pygocentrus_nattereri |
| ENSG00000101811 | CSTF2 | 96 | 98.140 | ENSRNOG00000048025 | Cstf2 | 100 | 92.759 | Rattus_norvegicus |
| ENSG00000101811 | CSTF2 | 100 | 97.920 | ENSRBIG00000030745 | CSTF2 | 100 | 97.920 | Rhinopithecus_bieti |
| ENSG00000101811 | CSTF2 | 100 | 99.665 | ENSRROG00000045042 | CSTF2 | 100 | 99.665 | Rhinopithecus_roxellana |
| ENSG00000101811 | CSTF2 | 100 | 98.995 | ENSSBOG00000021649 | CSTF2 | 100 | 98.995 | Saimiri_boliviensis_boliviensis |
| ENSG00000101811 | CSTF2 | 98 | 100.000 | ENSSHAG00000003681 | CSTF2 | 92 | 83.305 | Sarcophilus_harrisii |
| ENSG00000101811 | CSTF2 | 81 | 91.803 | ENSSFOG00015006348 | cstf2 | 98 | 63.471 | Scleropages_formosus |
| ENSG00000101811 | CSTF2 | 81 | 93.956 | ENSSMAG00000013463 | cstf2 | 97 | 59.354 | Scophthalmus_maximus |
| ENSG00000101811 | CSTF2 | 96 | 87.442 | ENSSPUG00000005288 | - | 98 | 80.720 | Sphenodon_punctatus |
| ENSG00000101811 | CSTF2 | 100 | 95.645 | ENSSSCG00000012486 | CSTF2 | 100 | 96.970 | Sus_scrofa |
| ENSG00000101811 | CSTF2 | 100 | 79.422 | ENSTGUG00000002253 | CSTF2 | 100 | 79.592 | Taeniopygia_guttata |
| ENSG00000101811 | CSTF2 | 83 | 89.785 | ENSTNIG00000016860 | cstf2 | 99 | 59.593 | Tetraodon_nigroviridis |
| ENSG00000101811 | CSTF2 | 55 | 100.000 | ENSTBEG00000010029 | - | 69 | 100.000 | Tupaia_belangeri |
| ENSG00000101811 | CSTF2 | 100 | 96.880 | ENSTTRG00000013556 | CSTF2 | 100 | 96.880 | Tursiops_truncatus |
| ENSG00000101811 | CSTF2 | 100 | 96.346 | ENSUAMG00000026605 | CSTF2 | 100 | 96.346 | Ursus_americanus |
| ENSG00000101811 | CSTF2 | 96 | 98.605 | ENSUMAG00000012417 | CSTF2 | 100 | 98.270 | Ursus_maritimus |
| ENSG00000101811 | CSTF2 | 95 | 98.598 | ENSVPAG00000006466 | CSTF2 | 72 | 96.210 | Vicugna_pacos |
| ENSG00000101811 | CSTF2 | 100 | 96.534 | ENSVVUG00000004717 | CSTF2 | 100 | 96.534 | Vulpes_vulpes |
| ENSG00000101811 | CSTF2 | 92 | 86.893 | ENSXETG00000009642 | cstf2 | 100 | 66.154 | Xenopus_tropicalis |
| ENSG00000101811 | CSTF2 | 75 | 94.675 | ENSXCOG00000016436 | cstf2 | 84 | 96.970 | Xiphophorus_couchianus |
| ENSG00000101811 | CSTF2 | 81 | 91.257 | ENSXMAG00000013524 | cstf2 | 97 | 56.070 | Xiphophorus_maculatus |
| Go ID | Go_term | PubmedID | Evidence | Category |
|---|---|---|---|---|
| GO:0000398 | mRNA splicing, via spliceosome | - | TAS | Process |
| GO:0003723 | RNA binding | 22658674.22681889. | HDA | Function |
| GO:0003723 | RNA binding | 21873635. | IBA | Function |
| GO:0003723 | RNA binding | 1741396. | IDA | Function |
| GO:0003729 | mRNA binding | 21873635. | IBA | Function |
| GO:0005515 | protein binding | 10477523.11598190.14707147.16189514.18688255.19136632.20379136.25416956. | IPI | Function |
| GO:0005654 | nucleoplasm | - | IDA | Component |
| GO:0005654 | nucleoplasm | - | TAS | Component |
| GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex | 21873635. | IBA | Component |
| GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex | 18305108.21102410. | IDA | Component |
| GO:0006369 | termination of RNA polymerase II transcription | - | TAS | Process |
| GO:0006378 | mRNA polyadenylation | - | IEA | Process |
| GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation | - | TAS | Process |
| GO:0016604 | nuclear body | - | IDA | Component |
| GO:0031124 | mRNA 3'-end processing | - | TAS | Process |
| GO:0071920 | cleavage body | 11598190. | IDA | Component |
| GO:0098789 | pre-mRNA cleavage required for polyadenylation | 21873635. | IBA | Process |