
| Cancer | Chr | Position | Mutation Type | dbSNP | Protein-change | Allele Freq | RBD |
|---|---|---|---|---|---|---|---|
| ACC | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CHOL | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| DLBC | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| KIRP | |||||||
| KIRP | |||||||
| LIHC | |||||||
| LIHC | |||||||
| LIHC | |||||||
| LIHC | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| OV | |||||||
| OV | |||||||
| OV | |||||||
| OV | |||||||
| OV | |||||||
| OV | |||||||
| PAAD | |||||||
| PAAD | |||||||
| PRAD | |||||||
| READ | |||||||
| READ | |||||||
| READ | |||||||
| READ | |||||||
| READ | |||||||
| SARC | |||||||
| SARC | |||||||
| SARC | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| THCA | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCS |
| Cancer | Type | Freq | Q-value |
|---|---|---|---|
| KIRP | |||
| LUAD | |||
| MESO |
| Cancer | P-value | Q-value |
|---|---|---|
| KIRC | ||
| LUSC | ||
| PAAD | ||
| LAML | ||
| GBM | ||
| DLBC | ||
| LUAD |
| Protein ID | Domain | Pfam ID | E-value | Domain number | Total number |
|---|---|---|---|---|---|
| ENSP00000281156 | KH_1 | PF00013.29 | 2.2e-08 | 1 | 1 |
| Transcript ID | Name | Length | RefSeq ID | Protein ID | Length | RefSeq ID | UniportKB ID |
|---|---|---|---|---|---|---|---|
| ENST00000281156 | KHDRBS2-201 | 2332 | XM_011535516 | ENSP00000281156 | 349 (aa) | XP_011533818 | Q5VWX1 |
| ensgID | Trait | pValue | Pubmed ID |
|---|---|---|---|
| ENSG00000112232 | Respiratory Function Tests | 5.25469189570592E-5 | 17903307 |
| ENSG00000112232 | Body Weights and Measures | 1.99036039627727E-6 | 17903300 |
| ENSG00000112232 | Cholesterol, HDL | 3.16857085789422E-5 | 17903299 |
| ENSG00000112232 | Cholesterol, HDL | 1.16519772628276E-5 | 17903299 |
| ENSG00000112232 | Cholesterol, HDL | 1.34720293036943E-8 | 17903299 |
| ENSG00000112232 | Uric Acid | 8.78169922178608E-5 | 17903292 |
| ENSG00000112232 | Alcohol Drinking | 2.26327203259413E-10 | - |
| ENSG00000112232 | Pulse | 1.0066776280682E-5 | 17903302 |
| ENSG00000112232 | Pulse | 1.4808784910726E-5 | 17903302 |
| ENSG00000112232 | Arteries | 5.8810000E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 4.8112592E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.6083107E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.8668589E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 3.3564102E-006 | - |
| ENSG00000112232 | Leukoaraiosis | 3.3576568E-006 | - |
| ENSG00000112232 | Leukoaraiosis | 1.8227434E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 2.2220383E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.7901518E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 4.6955880E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 3.4838039E-006 | - |
| ENSG00000112232 | Leukoaraiosis | 8.7323639E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 9.7966349E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 9.6918656E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 7.7200054E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 7.2344517E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.7621139E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.8702547E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.8439355E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.6929706E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.7318140E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.6720066E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.3152022E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.2941446E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 6.8820744E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 1.6289055E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 6.9991765E-006 | - |
| ENSG00000112232 | Leukoaraiosis | 1.9601693E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 1.5778443E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 7.0038028E-006 | - |
| ENSG00000112232 | Leukoaraiosis | 3.6109710E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 8.8026608E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 6.6673340E-006 | - |
| ENSG00000112232 | Leukoaraiosis | 3.1215111E-005 | - |
| ENSG00000112232 | Leukoaraiosis | 6.6721126E-006 | - |
| ENSG00000112232 | Leukoaraiosis | 8.2306231E-006 | - |
| ENSG00000112232 | Leukoaraiosis | 8.7232461E-005 | - |
| ENSG00000112232 | Anatomy | 7.3065816E-005 | - |
| ENSG00000112232 | Anatomy | 6.9735737E-005 | - |
| ENSG00000112232 | Platelet Function Tests | 3.8137661E-007 | - |
| ENSG00000112232 | Platelet Function Tests | 7.2271080E-012 | - |
| ENSG00000112232 | Platelet Function Tests | 1.2141200E-005 | - |
| ENSG00000112232 | Platelet Function Tests | 2.1322200E-005 | - |
| ENSG00000112232 | Platelet Function Tests | 1.7635400E-005 | - |
| ENSG00000112232 | Platelet Function Tests | 2.1325300E-005 | - |
| ENSG00000112232 | Platelet Function Tests | 1.2141200E-005 | - |
| ENSG00000112232 | Platelet Function Tests | 6.0122759E-006 | - |
| ENSG00000112232 | Platelet Function Tests | 4.9823613E-006 | - |
| ENSG00000112232 | Platelet Function Tests | 6.0122759E-006 | - |
| ENSG00000112232 | Platelet Function Tests | 6.5795716E-006 | - |
| ENSG00000112232 | Blood Pressure | 2.6626382E-010 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 5.4490000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 6.4900000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 6.3010000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 5.0380000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 3.0520000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 1.2420000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 4.8000000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 3.7160000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 1.4220000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 3.4260000E-006 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 6.7850000E-006 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 4.7230000E-006 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 1.5190000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 5.4540000E-006 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 1.7710000E-005 | - |
| ENSG00000112232 | Aspartate Aminotransferases | 3.4700000E-006 | - |
| ENSG00000112232 | Alzheimer Disease | 9E-7 | 26830138 |
| ENSG00000112232 | Age of Onset | 9E-7 | 26830138 |
| ENSG00000112232 | Adenocarcinoma Of Esophagus | 2E-8 | 27527254 |
| ENSG00000112232 | Hip | 5E-6 | 23251661 |
| ENSG00000112232 | Antibodies, Antiphospholipid | 4E-6 | 27098658 |
| ENSG00000112232 | Aspartate Aminotransferases | 3E-7 | 18464913 |
| ENSG00000112232 | Calcium | 3E-6 | 20705733 |
| ensgID | SNP | Chromosome | Position | SNP-risk | Trait | PubmedID | 95% CI | Or or BEAT | EFO ID |
|---|---|---|---|---|---|---|---|---|---|
| ENSG00000112232 | rs34540100 | 6 | 62264103 | ? | Presence of antiphospholipid antibodies | 27098658 | EFO_0005200 | ||
| ENSG00000112232 | rs76014404 | 6 | 61681634 | T | Esophageal adenocarcinoma | 27527254 | [1.14-1.28] | 1.2102 | EFO_0000478 |
| ENSG00000112232 | rs11324540 | 6 | 62002189 | ? | Epstein-Barr virus copy number in lymphoblastoid cell lines | 28654678 | EFO_0000769 | ||
| ENSG00000112232 | rs855408 | 6 | 62257812 | ? | Epstein-Barr virus copy number in lymphoblastoid cell lines | 28654678 | EFO_0000769 | ||
| ENSG00000112232 | rs9363882 | 6 | 62150634 | T | Antipsychotic drug-induced QTc interval change in schizophrenia | 29064910 | unit decrease | 5350 | EFO_0000692|EFO_0008003|GO_0097332 |
| ENSG00000112232 | rs998808 | 6 | 61997852 | A | Antipsychotic drug-induced QTc interval change in schizophrenia | 29064910 | unit decrease | 6910 | EFO_0000692|EFO_0008003|GO_0097332 |
| ENSG00000112232 | rs6908169 | 6 | 61991720 | C | Self-reported math ability | 30038396 | [0.0072-0.015] unit increase | 0.0111 | EFO_0004875 |
| ENSG00000112232 | rs200082850 | 6 | 62240663 | ATG | beta-nerve growth factor levels | 27989323 | [0.086-0.187] SD units decrease | 0.1363 | EFO_0008035 |
| ENSG00000112232 | rs6455128 | 6 | 61987841 | ? | Protein quantitative trait loci | 18464913 | EFO_0004736 | ||
| ENSG00000112232 | rs1931814 | 6 | 61879262 | ? | Morning person | 30595370 | EFO_0008328 | ||
| ENSG00000112232 | rs12524852 | 6 | 61973218 | ? | Sunburns | 30595370 | EFO_0003958 | ||
| ENSG00000112232 | rs59899220 | 6 | 61822220 | ? | Lung function (FEV1/FVC) | 30595370 | EFO_0004713 | ||
| ENSG00000112232 | rs10223780 | 6 | 62155120 | G | Corneal astigmatism | 30306274 | [1.035-1.083] | 1.059 | EFO_1002040 |
| ENSG00000112232 | rs1931814 | 6 | 61879262 | A | Chronotype | 30696823 | 1.0267515 | EFO_0008328 |
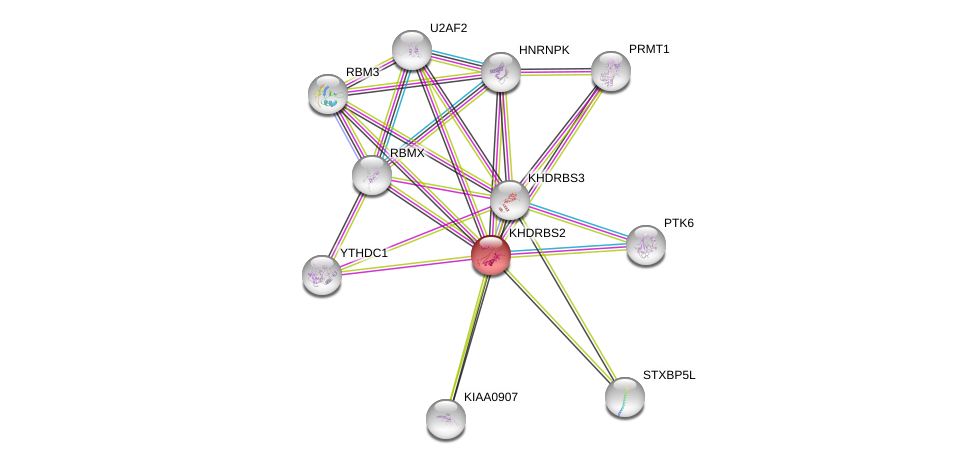
| Ensembl ID | Gene Symbol | Coverage | Identiy | Paralog | Gene Symbol | Coverage | Identiy |
|---|---|---|---|---|---|---|---|
| ENSG00000112232 | KHDRBS2 | 99 | 65.730 | ENSG00000131773 | KHDRBS3 | 94 | 79.221 |
| ENSG00000112232 | KHDRBS2 | 99 | 60.773 | ENSG00000121774 | KHDRBS1 | 78 | 62.040 |
| ENSG00000112232 | KHDRBS2 | 77 | 36.396 | ENSG00000112531 | QKI | 76 | 55.128 |
| Ensembl ID | Gene Symbol | Coverage | Identiy | Ortholog | Gene Symbol | Coverage | Identiy | Species |
|---|---|---|---|---|---|---|---|---|
| ENSG00000112232 | KHDRBS2 | 100 | 69.034 | ENSAPOG00000008806 | khdrbs2 | 100 | 68.466 | Acanthochromis_polyacanthus |
| ENSG00000112232 | KHDRBS2 | 77 | 98.889 | ENSAMEG00000007111 | KHDRBS2 | 100 | 98.889 | Ailuropoda_melanoleuca |
| ENSG00000112232 | KHDRBS2 | 100 | 68.466 | ENSACIG00000020525 | khdrbs2 | 100 | 67.705 | Amphilophus_citrinellus |
| ENSG00000112232 | KHDRBS2 | 100 | 67.898 | ENSAOCG00000011333 | khdrbs2 | 100 | 65.909 | Amphiprion_ocellaris |
| ENSG00000112232 | KHDRBS2 | 95 | 65.476 | ENSAPEG00000019843 | khdrbs2 | 99 | 64.478 | Amphiprion_percula |
| ENSG00000112232 | KHDRBS2 | 100 | 68.750 | ENSATEG00000020863 | khdrbs2 | 100 | 67.806 | Anabas_testudineus |
| ENSG00000112232 | KHDRBS2 | 100 | 93.410 | ENSAPLG00000013452 | KHDRBS2 | 100 | 93.410 | Anas_platyrhynchos |
| ENSG00000112232 | KHDRBS2 | 100 | 89.971 | ENSACAG00000006691 | KHDRBS2 | 100 | 88.825 | Anolis_carolinensis |
| ENSG00000112232 | KHDRBS2 | 85 | 98.990 | ENSANAG00000024792 | KHDRBS2 | 99 | 98.990 | Aotus_nancymaae |
| ENSG00000112232 | KHDRBS2 | 100 | 68.555 | ENSACLG00000002710 | khdrbs2 | 100 | 67.989 | Astatotilapia_calliptera |
| ENSG00000112232 | KHDRBS2 | 88 | 74.434 | ENSAMXG00000009582 | khdrbs2 | 89 | 74.434 | Astyanax_mexicanus |
| ENSG00000112232 | KHDRBS2 | 100 | 92.837 | ENSBTAG00000043990 | KHDRBS2 | 100 | 93.983 | Bos_taurus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.281 | ENSCJAG00000001766 | KHDRBS2 | 100 | 98.281 | Callithrix_jacchus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.281 | ENSCAFG00000002470 | KHDRBS2 | 100 | 98.281 | Canis_familiaris |
| ENSG00000112232 | KHDRBS2 | 100 | 98.281 | ENSCAFG00020015926 | KHDRBS2 | 100 | 98.281 | Canis_lupus_dingo |
| ENSG00000112232 | KHDRBS2 | 100 | 92.550 | ENSCHIG00000024242 | KHDRBS2 | 100 | 92.550 | Capra_hircus |
| ENSG00000112232 | KHDRBS2 | 91 | 99.057 | ENSTSYG00000007203 | KHDRBS2 | 91 | 99.057 | Carlito_syrichta |
| ENSG00000112232 | KHDRBS2 | 91 | 94.969 | ENSCPOG00000025621 | KHDRBS2 | 100 | 94.969 | Cavia_porcellus |
| ENSG00000112232 | KHDRBS2 | 100 | 85.960 | ENSCCAG00000037746 | KHDRBS2 | 100 | 85.960 | Cebus_capucinus |
| ENSG00000112232 | KHDRBS2 | 100 | 99.427 | ENSCATG00000042976 | KHDRBS2 | 100 | 99.427 | Cercocebus_atys |
| ENSG00000112232 | KHDRBS2 | 91 | 96.541 | ENSCLAG00000000509 | KHDRBS2 | 100 | 96.541 | Chinchilla_lanigera |
| ENSG00000112232 | KHDRBS2 | 77 | 100.000 | ENSCSAG00000013462 | KHDRBS2 | 98 | 100.000 | Chlorocebus_sabaeus |
| ENSG00000112232 | KHDRBS2 | 100 | 69.054 | ENSCHOG00000013616 | KHDRBS2 | 100 | 68.768 | Choloepus_hoffmanni |
| ENSG00000112232 | KHDRBS2 | 100 | 93.410 | ENSCPBG00000022649 | KHDRBS2 | 100 | 93.410 | Chrysemys_picta_bellii |
| ENSG00000112232 | KHDRBS2 | 100 | 99.427 | ENSCANG00000021348 | KHDRBS2 | 100 | 99.427 | Colobus_angolensis_palliatus |
| ENSG00000112232 | KHDRBS2 | 100 | 95.702 | ENSCGRG00001022657 | Khdrbs2 | 100 | 95.702 | Cricetulus_griseus_chok1gshd |
| ENSG00000112232 | KHDRBS2 | 77 | 96.296 | ENSCGRG00000003079 | - | 96 | 96.296 | Cricetulus_griseus_crigri |
| ENSG00000112232 | KHDRBS2 | 100 | 58.739 | ENSCSEG00000005064 | khdrbs2 | 100 | 62.286 | Cynoglossus_semilaevis |
| ENSG00000112232 | KHDRBS2 | 77 | 85.106 | ENSCVAG00000020560 | khdrbs2 | 100 | 59.312 | Cyprinodon_variegatus |
| ENSG00000112232 | KHDRBS2 | 100 | 72.571 | ENSDARG00000069469 | khdrbs2 | 100 | 72.286 | Danio_rerio |
| ENSG00000112232 | KHDRBS2 | 100 | 96.848 | ENSDNOG00000014028 | KHDRBS2 | 100 | 96.848 | Dasypus_novemcinctus |
| ENSG00000112232 | KHDRBS2 | 94 | 87.462 | ENSDORG00000005701 | Khdrbs2 | 100 | 88.679 | Dipodomys_ordii |
| ENSG00000112232 | KHDRBS2 | 100 | 79.370 | ENSETEG00000002241 | KHDRBS2 | 100 | 79.370 | Echinops_telfairi |
| ENSG00000112232 | KHDRBS2 | 93 | 56.395 | ENSEBUG00000007730 | khdrbs2 | 100 | 51.902 | Eptatretus_burgeri |
| ENSG00000112232 | KHDRBS2 | 79 | 47.973 | ENSEBUG00000012421 | khdrbs2 | 99 | 50.336 | Eptatretus_burgeri |
| ENSG00000112232 | KHDRBS2 | 99 | 47.965 | ENSEBUG00000007596 | khdrbs2 | 88 | 50.145 | Eptatretus_burgeri |
| ENSG00000112232 | KHDRBS2 | 100 | 98.854 | ENSEASG00005003578 | KHDRBS2 | 100 | 98.854 | Equus_asinus_asinus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.567 | ENSECAG00000021541 | KHDRBS2 | 100 | 98.567 | Equus_caballus |
| ENSG00000112232 | KHDRBS2 | 91 | 96.845 | ENSEEUG00000003986 | KHDRBS2 | 100 | 96.845 | Erinaceus_europaeus |
| ENSG00000112232 | KHDRBS2 | 100 | 72.080 | ENSELUG00000020898 | khdrbs2 | 100 | 69.429 | Esox_lucius |
| ENSG00000112232 | KHDRBS2 | 100 | 98.567 | ENSFCAG00000046207 | KHDRBS2 | 78 | 98.567 | Felis_catus |
| ENSG00000112232 | KHDRBS2 | 91 | 91.223 | ENSFALG00000001691 | KHDRBS2 | 99 | 96.450 | Ficedula_albicollis |
| ENSG00000112232 | KHDRBS2 | 91 | 85.220 | ENSFDAG00000008941 | KHDRBS2 | 100 | 87.107 | Fukomys_damarensis |
| ENSG00000112232 | KHDRBS2 | 81 | 49.310 | ENSFHEG00000021135 | khdrbs2 | 89 | 47.635 | Fundulus_heteroclitus |
| ENSG00000112232 | KHDRBS2 | 100 | 62.040 | ENSGMOG00000005065 | khdrbs2 | 100 | 61.972 | Gadus_morhua |
| ENSG00000112232 | KHDRBS2 | 100 | 92.550 | ENSGALG00000016276 | KHDRBS2 | 100 | 91.117 | Gallus_gallus |
| ENSG00000112232 | KHDRBS2 | 100 | 67.806 | ENSGAFG00000007645 | khdrbs2 | 100 | 66.761 | Gambusia_affinis |
| ENSG00000112232 | KHDRBS2 | 100 | 67.705 | ENSGACG00000006097 | khdrbs2 | 100 | 67.045 | Gasterosteus_aculeatus |
| ENSG00000112232 | KHDRBS2 | 58 | 91.089 | ENSGAGG00000023457 | - | 71 | 89.109 | Gopherus_agassizii |
| ENSG00000112232 | KHDRBS2 | 100 | 100.000 | ENSGGOG00000025068 | KHDRBS2 | 100 | 100.000 | Gorilla_gorilla |
| ENSG00000112232 | KHDRBS2 | 100 | 68.555 | ENSHBUG00000008029 | khdrbs2 | 100 | 67.989 | Haplochromis_burtoni |
| ENSG00000112232 | KHDRBS2 | 100 | 94.556 | ENSHGLG00000017439 | KHDRBS2 | 100 | 94.556 | Heterocephalus_glaber_female |
| ENSG00000112232 | KHDRBS2 | 100 | 94.556 | ENSHGLG00100003817 | KHDRBS2 | 100 | 94.556 | Heterocephalus_glaber_male |
| ENSG00000112232 | KHDRBS2 | 100 | 62.323 | ENSHCOG00000013857 | khdrbs2 | 100 | 62.040 | Hippocampus_comes |
| ENSG00000112232 | KHDRBS2 | 100 | 74.286 | ENSIPUG00000015437 | khdrbs2 | 100 | 74.286 | Ictalurus_punctatus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.854 | ENSSTOG00000011529 | KHDRBS2 | 100 | 98.854 | Ictidomys_tridecemlineatus |
| ENSG00000112232 | KHDRBS2 | 100 | 96.562 | ENSJJAG00000017812 | Khdrbs2 | 100 | 96.562 | Jaculus_jaculus |
| ENSG00000112232 | KHDRBS2 | 100 | 69.318 | ENSKMAG00000018214 | khdrbs2 | 100 | 68.750 | Kryptolebias_marmoratus |
| ENSG00000112232 | KHDRBS2 | 99 | 66.762 | ENSLBEG00000013077 | khdrbs2 | 86 | 67.529 | Labrus_bergylta |
| ENSG00000112232 | KHDRBS2 | 100 | 79.714 | ENSLOCG00000017016 | khdrbs2 | 98 | 79.714 | Lepisosteus_oculatus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.567 | ENSLAFG00000006655 | KHDRBS2 | 100 | 98.567 | Loxodonta_africana |
| ENSG00000112232 | KHDRBS2 | 100 | 99.140 | ENSMFAG00000002823 | KHDRBS2 | 100 | 99.140 | Macaca_fascicularis |
| ENSG00000112232 | KHDRBS2 | 100 | 99.427 | ENSMNEG00000042875 | KHDRBS2 | 100 | 99.427 | Macaca_nemestrina |
| ENSG00000112232 | KHDRBS2 | 77 | 100.000 | ENSMLEG00000022013 | KHDRBS2 | 100 | 100.000 | Mandrillus_leucophaeus |
| ENSG00000112232 | KHDRBS2 | 100 | 68.091 | ENSMAMG00000015552 | khdrbs2 | 100 | 68.376 | Mastacembelus_armatus |
| ENSG00000112232 | KHDRBS2 | 100 | 67.989 | ENSMZEG00005005504 | khdrbs2 | 100 | 67.422 | Maylandia_zebra |
| ENSG00000112232 | KHDRBS2 | 56 | 94.416 | ENSMAUG00000013631 | Khdrbs2 | 82 | 82.328 | Mesocricetus_auratus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.281 | ENSMICG00000010315 | KHDRBS2 | 100 | 98.281 | Microcebus_murinus |
| ENSG00000112232 | KHDRBS2 | 100 | 94.269 | ENSMOCG00000012101 | Khdrbs2 | 100 | 94.269 | Microtus_ochrogaster |
| ENSG00000112232 | KHDRBS2 | 100 | 67.330 | ENSMMOG00000007708 | khdrbs2 | 100 | 65.909 | Mola_mola |
| ENSG00000112232 | KHDRBS2 | 100 | 93.696 | ENSMODG00000018714 | KHDRBS2 | 100 | 93.696 | Monodelphis_domestica |
| ENSG00000112232 | KHDRBS2 | 100 | 65.909 | ENSMALG00000002214 | khdrbs2 | 100 | 65.625 | Monopterus_albus |
| ENSG00000112232 | KHDRBS2 | 100 | 94.842 | MGP_CAROLIEiJ_G0014010 | Khdrbs2 | 100 | 94.842 | Mus_caroli |
| ENSG00000112232 | KHDRBS2 | 100 | 95.415 | ENSMUSG00000026058 | Khdrbs2 | 100 | 95.415 | Mus_musculus |
| ENSG00000112232 | KHDRBS2 | 100 | 95.129 | MGP_PahariEiJ_G0027259 | Khdrbs2 | 100 | 95.129 | Mus_pahari |
| ENSG00000112232 | KHDRBS2 | 100 | 95.415 | MGP_SPRETEiJ_G0014814 | Khdrbs2 | 100 | 95.415 | Mus_spretus |
| ENSG00000112232 | KHDRBS2 | 100 | 99.140 | ENSMPUG00000002607 | KHDRBS2 | 100 | 99.140 | Mustela_putorius_furo |
| ENSG00000112232 | KHDRBS2 | 100 | 97.994 | ENSMLUG00000005484 | KHDRBS2 | 100 | 97.994 | Myotis_lucifugus |
| ENSG00000112232 | KHDRBS2 | 100 | 68.555 | ENSNBRG00000013759 | khdrbs2 | 100 | 67.989 | Neolamprologus_brichardi |
| ENSG00000112232 | KHDRBS2 | 91 | 94.340 | ENSODEG00000015927 | KHDRBS2 | 100 | 94.340 | Octodon_degus |
| ENSG00000112232 | KHDRBS2 | 100 | 68.927 | ENSONIG00000009755 | khdrbs2 | 100 | 67.989 | Oreochromis_niloticus |
| ENSG00000112232 | KHDRBS2 | 100 | 96.848 | ENSOCUG00000012344 | KHDRBS2 | 100 | 96.848 | Oryctolagus_cuniculus |
| ENSG00000112232 | KHDRBS2 | 100 | 67.614 | ENSORLG00000010165 | khdrbs2 | 100 | 67.330 | Oryzias_latipes |
| ENSG00000112232 | KHDRBS2 | 100 | 67.330 | ENSORLG00020020168 | khdrbs2 | 100 | 66.761 | Oryzias_latipes_hni |
| ENSG00000112232 | KHDRBS2 | 100 | 67.330 | ENSORLG00015003971 | khdrbs2 | 100 | 66.761 | Oryzias_latipes_hsok |
| ENSG00000112232 | KHDRBS2 | 100 | 67.045 | ENSOMEG00000001684 | khdrbs2 | 100 | 66.477 | Oryzias_melastigma |
| ENSG00000112232 | KHDRBS2 | 100 | 98.281 | ENSOGAG00000015071 | KHDRBS2 | 100 | 98.281 | Otolemur_garnettii |
| ENSG00000112232 | KHDRBS2 | 95 | 91.124 | ENSOARG00000005603 | KHDRBS2 | 97 | 91.124 | Ovis_aries |
| ENSG00000112232 | KHDRBS2 | 100 | 99.713 | ENSPPAG00000032651 | KHDRBS2 | 100 | 99.713 | Pan_paniscus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.854 | ENSPPRG00000008147 | KHDRBS2 | 100 | 98.854 | Panthera_pardus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.854 | ENSPTIG00000020547 | KHDRBS2 | 100 | 98.854 | Panthera_tigris_altaica |
| ENSG00000112232 | KHDRBS2 | 100 | 100.000 | ENSPTRG00000018311 | KHDRBS2 | 100 | 100.000 | Pan_troglodytes |
| ENSG00000112232 | KHDRBS2 | 100 | 99.427 | ENSPANG00000021881 | KHDRBS2 | 100 | 99.427 | Papio_anubis |
| ENSG00000112232 | KHDRBS2 | 100 | 68.946 | ENSPMGG00000008580 | khdrbs2 | 100 | 68.661 | Periophthalmus_magnuspinnatus |
| ENSG00000112232 | KHDRBS2 | 100 | 68.272 | ENSPFOG00000011638 | khdrbs2 | 100 | 67.330 | Poecilia_formosa |
| ENSG00000112232 | KHDRBS2 | 100 | 66.952 | ENSPLAG00000023478 | khdrbs2 | 100 | 66.193 | Poecilia_latipinna |
| ENSG00000112232 | KHDRBS2 | 100 | 67.521 | ENSPMEG00000011563 | khdrbs2 | 100 | 66.761 | Poecilia_mexicana |
| ENSG00000112232 | KHDRBS2 | 100 | 67.898 | ENSPREG00000022525 | khdrbs2 | 100 | 67.330 | Poecilia_reticulata |
| ENSG00000112232 | KHDRBS2 | 58 | 100.000 | ENSPPYG00000029598 | - | 97 | 100.000 | Pongo_abelii |
| ENSG00000112232 | KHDRBS2 | 100 | 84.814 | ENSPCAG00000001426 | KHDRBS2 | 100 | 84.814 | Procavia_capensis |
| ENSG00000112232 | KHDRBS2 | 100 | 85.387 | ENSPCOG00000018881 | KHDRBS2 | 100 | 85.387 | Propithecus_coquereli |
| ENSG00000112232 | KHDRBS2 | 100 | 68.555 | ENSPNYG00000019550 | khdrbs2 | 100 | 67.989 | Pundamilia_nyererei |
| ENSG00000112232 | KHDRBS2 | 99 | 75.575 | ENSPNAG00000008598 | khdrbs2 | 99 | 75.575 | Pygocentrus_nattereri |
| ENSG00000112232 | KHDRBS2 | 100 | 95.415 | ENSRNOG00000012284 | Khdrbs2 | 100 | 95.415 | Rattus_norvegicus |
| ENSG00000112232 | KHDRBS2 | 91 | 98.734 | ENSRBIG00000039774 | KHDRBS2 | 99 | 98.734 | Rhinopithecus_bieti |
| ENSG00000112232 | KHDRBS2 | 100 | 97.994 | ENSSBOG00000019779 | KHDRBS2 | 100 | 97.994 | Saimiri_boliviensis_boliviensis |
| ENSG00000112232 | KHDRBS2 | 100 | 76.286 | ENSSFOG00015012491 | khdrbs2 | 100 | 75.429 | Scleropages_formosus |
| ENSG00000112232 | KHDRBS2 | 99 | 66.856 | ENSSMAG00000006422 | khdrbs2 | 89 | 66.572 | Scophthalmus_maximus |
| ENSG00000112232 | KHDRBS2 | 100 | 70.085 | ENSSDUG00000023520 | khdrbs2 | 100 | 69.516 | Seriola_dumerili |
| ENSG00000112232 | KHDRBS2 | 99 | 69.429 | ENSSLDG00000019363 | khdrbs2 | 97 | 69.253 | Seriola_lalandi_dorsalis |
| ENSG00000112232 | KHDRBS2 | 100 | 75.645 | ENSSARG00000004121 | KHDRBS2 | 100 | 75.645 | Sorex_araneus |
| ENSG00000112232 | KHDRBS2 | 70 | 92.593 | ENSSPUG00000006155 | KHDRBS2 | 77 | 92.593 | Sphenodon_punctatus |
| ENSG00000112232 | KHDRBS2 | 100 | 97.994 | ENSSSCG00000001490 | KHDRBS2 | 100 | 97.994 | Sus_scrofa |
| ENSG00000112232 | KHDRBS2 | 69 | 92.500 | ENSTGUG00000012786 | KHDRBS2 | 99 | 90.417 | Taeniopygia_guttata |
| ENSG00000112232 | KHDRBS2 | 100 | 65.156 | ENSTRUG00000015760 | khdrbs2 | 100 | 67.045 | Takifugu_rubripes |
| ENSG00000112232 | KHDRBS2 | 100 | 96.275 | ENSTTRG00000005591 | KHDRBS2 | 100 | 96.275 | Tursiops_truncatus |
| ENSG00000112232 | KHDRBS2 | 100 | 98.854 | ENSUMAG00000018593 | KHDRBS2 | 100 | 98.854 | Ursus_maritimus |
| ENSG00000112232 | KHDRBS2 | 91 | 96.215 | ENSVPAG00000001469 | KHDRBS2 | 100 | 96.215 | Vicugna_pacos |
| ENSG00000112232 | KHDRBS2 | 100 | 98.281 | ENSVVUG00000008652 | KHDRBS2 | 100 | 98.281 | Vulpes_vulpes |
| ENSG00000112232 | KHDRBS2 | 100 | 77.650 | ENSXETG00000004451 | khdrbs2 | 100 | 74.499 | Xenopus_tropicalis |
| ENSG00000112232 | KHDRBS2 | 100 | 68.466 | ENSXMAG00000009415 | khdrbs2 | 100 | 66.761 | Xiphophorus_maculatus |
| Go ID | Go_term | PubmedID | Evidence | Category |
|---|---|---|---|---|
| GO:0005515 | protein binding | 16189514.19282290.21516116.25416956.25910212.26871637.27107012. | IPI | Function |
| GO:0005654 | nucleoplasm | - | TAS | Component |
| GO:0006397 | mRNA processing | - | IEA | Process |
| GO:0008143 | poly(A) binding | - | IEA | Function |
| GO:0008266 | poly(U) RNA binding | - | IEA | Function |
| GO:0017124 | SH3 domain binding | - | IEA | Function |
| GO:0042169 | SH2 domain binding | - | IEA | Function |
| GO:0042802 | identical protein binding | 21516116.25416956. | IPI | Function |
| GO:0046982 | protein heterodimerization activity | - | IEA | Function |
| GO:0048024 | regulation of mRNA splicing, via spliceosome | - | IEA | Process |