
| Cancer | Chr | Position | Mutation Type | dbSNP | Protein-change | Allele Freq | RBD |
|---|---|---|---|---|---|---|---|
| ACC | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| CHOL | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| ESCA | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| HNSC | |||||||
| KIRC | |||||||
| LGG | |||||||
| LGG | |||||||
| LIHC | |||||||
| LIHC | |||||||
| LIHC | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| OV | |||||||
| PAAD | |||||||
| PRAD | |||||||
| PRAD | |||||||
| PRAD | |||||||
| PRAD | |||||||
| PRAD | |||||||
| READ | |||||||
| READ | |||||||
| READ | |||||||
| SARC | |||||||
| SARC | |||||||
| SARC | |||||||
| SARC | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| THYM | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCS |
| Cancer | Type | Freq | Q-value |
|---|---|---|---|
| KIRC | |||
| PRAD | |||
| READ | |||
| SARC | |||
| TGCT | |||
| THCA |
| Cancer | P-value | Q-value |
|---|---|---|
| KIRC | ||
| STAD | ||
| UCS | ||
| SKCM | ||
| PAAD | ||
| PCPG | ||
| KICH | ||
| UCEC | ||
| LIHC | ||
| DLBC | ||
| LGG |
| Protein ID | Domain | Pfam ID | E-value | Domain number | Total number |
|---|---|---|---|---|---|
| ENSP00000427787 | RRM_1 | PF00076.22 | 1.3e-11 | 1 | 1 |
| ENSP00000428711 | RRM_1 | PF00076.22 | 1.3e-11 | 1 | 1 |
| ENSP00000430065 | RRM_1 | PF00076.22 | 1.2e-10 | 1 | 1 |
| ENSP00000428667 | RRM_1 | PF00076.22 | 1.2e-10 | 1 | 1 |
| ENSP00000430367 | RRM_1 | PF00076.22 | 1.2e-10 | 1 | 1 |
| ENSP00000430394 | RRM_1 | PF00076.22 | 1.2e-10 | 1 | 1 |
| ENSP00000430128 | RRM_1 | PF00076.22 | 1.3e-10 | 1 | 1 |
| PID | Title | Method | Time | Author | Doi |
|---|---|---|---|---|---|
| 22658674 |
| Transcript ID | Name | Length | RefSeq ID | Protein ID | Length | RefSeq ID | UniportKB ID |
|---|---|---|---|---|---|---|---|
| ENST00000521695 | RALYL-207 | 1924 | - | ENSP00000428667 | 291 (aa) | - | Q86SE5 |
| ENST00000521376 | RALYL-206 | 1457 | - | ENSP00000428310 | 146 (aa) | - | E5RJ39 |
| ENST00000523850 | RALYL-211 | 1684 | XM_017013084 | ENSP00000428807 | 218 (aa) | XP_016868573 | Q86SE5 |
| ENST00000521268 | RALYL-205 | 2701 | - | ENSP00000430367 | 291 (aa) | - | Q86SE5 |
| ENST00000522455 | RALYL-208 | 2168 | - | ENSP00000430394 | 291 (aa) | - | Q86SE5 |
| ENST00000518065 | RALYL-203 | 1505 | - | - | - (aa) | - | - |
| ENST00000517638 | RALYL-201 | 1972 | XM_024447069 | ENSP00000430128 | 304 (aa) | XP_024302837 | Q86SE5 |
| ENST00000517988 | RALYL-202 | 571 | - | ENSP00000428711 | 85 (aa) | - | E5RG71 |
| ENST00000522613 | RALYL-209 | 595 | - | ENSP00000427787 | 85 (aa) | - | E5RG71 |
| ENST00000518566 | RALYL-204 | 2682 | XM_011517462 | ENSP00000430065 | 280 (aa) | XP_011515764 | B3KT61 |
| ENST00000522647 | RALYL-210 | 459 | - | ENSP00000429284 | 47 (aa) | - | E5RIX9 |
| ensgID | Trait | pValue | Pubmed ID |
|---|---|---|---|
| ENSG00000184672 | Cholesterol, LDL | 1.44994943160715E-7 | 17903299 |
| ENSG00000184672 | Cholesterol, LDL | 1.32792993124453E-6 | 17903299 |
| ENSG00000184672 | Aorta | 3.73973094601978E-9 | 17903303 |
| ENSG00000184672 | Aorta | 9.79578646262135E-7 | 17903303 |
| ENSG00000184672 | Echocardiography | 1.90281222396393E-5 | 17903301 |
| ENSG00000184672 | Echocardiography | 6.46942529538563E-5 | 17903301 |
| ENSG00000184672 | Neuroblastoma | 2.79E-7 | 18463370 |
| ENSG00000184672 | Neuroblastoma | 1.08E-6 | 18463370 |
| ENSG00000184672 | Stroke | 5.2330859E-004 | 19369658 |
| ENSG00000184672 | Blood Pressure | 8.6030000E-005 | - |
| ENSG00000184672 | Cardiovascular Physiological Phenomena | 4.1510000E-005 | - |
| ENSG00000184672 | Coronary Disease | 2.8920000E-005 | - |
| ENSG00000184672 | Platelet Count | 9.8770000E-005 | - |
| ENSG00000184672 | Platelet Function Tests | 2.6804000E-005 | - |
| ENSG00000184672 | Platelet Function Tests | 3.0813900E-005 | - |
| ENSG00000184672 | Platelet Function Tests | 8.4700000E-006 | - |
| ENSG00000184672 | Platelet Function Tests | 1.2994300E-005 | - |
| ENSG00000184672 | Platelet Function Tests | 1.2994300E-005 | - |
| ENSG00000184672 | Cryptorchidism | 1.5200000E-016 | - |
| ENSG00000184672 | Sleep | 4E-6 | 27126917 |
| ensgID | SNP | Chromosome | Position | SNP-risk | Trait | PubmedID | 95% CI | Or or BEAT | EFO ID |
|---|---|---|---|---|---|---|---|---|---|
| ENSG00000184672 | rs118149821 | 8 | 84363832 | T | Night sleep phenotypes | 27126917 | [0.19-0.47] unit increase | 0.327 | EFO_0005272 |
| ENSG00000184672 | rs117893056 | 8 | 84374539 | T | Intelligence (MTAG) | 29326435 | [0.014-0.03] unit decrease | 0.022196304 | EFO_0004337 |
| ENSG00000184672 | rs113714691 | 8 | 84892347 | T | General cognitive ability | 29844566 | z-score decrease | 6.368 | EFO_0004337 |
| ENSG00000184672 | rs78565420 | 8 | 84790830 | T | General cognitive ability | 29844566 | z-score decrease | 5.559 | EFO_0004337 |
| ENSG00000184672 | rs783783 | 8 | 84491421 | ? | General cognitive ability | 29844566 | z-score decrease | 5.165 | EFO_0004337 |
| ENSG00000184672 | rs67589009 | 8 | 84326199 | ? | General cognitive ability | 29844566 | z-score increase | 5.085 | EFO_0004337 |
| ENSG00000184672 | rs2634047 | 8 | 84784102 | ? | Body mass index | 30595370 | EFO_0004340 | ||
| ENSG00000184672 | rs12677170 | 8 | 84534332 | C | Depressive symptoms x independent stressful life events interaction (1df test) | 30718454 | NR unit decrease | 0.074386 | EFO_0007006|EFO_0007781 |
| ENSG00000184672 | rs12677170 | 8 | 84534332 | C | Depressive symptoms x independent stressful life events interaction (2df test) | 30718454 | NR unit decrease | 0.0295941 | EFO_0007006|EFO_0007781 |
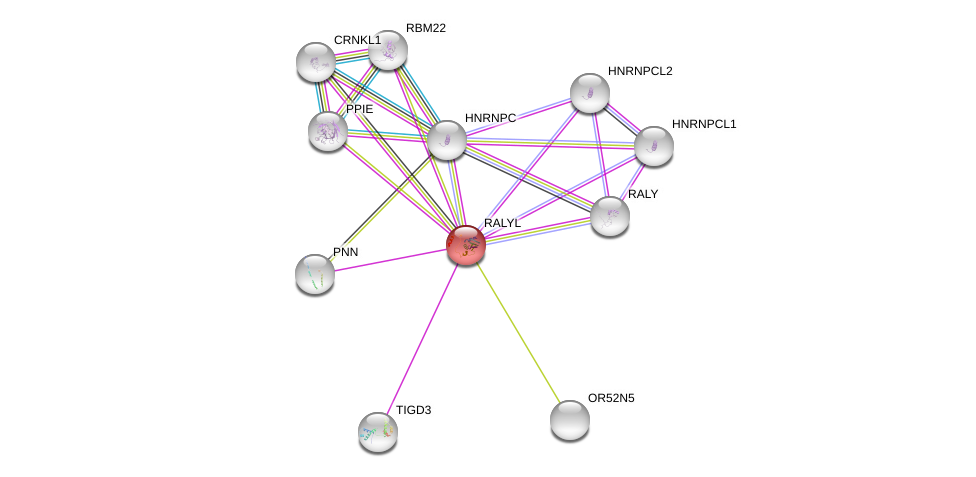
| Ensembl ID | Gene Symbol | Coverage | Identiy | Paralog | Gene Symbol | Coverage | Identiy |
|---|---|---|---|---|---|---|---|
| ENSG00000184672 | RALYL | 87 | 80.488 | ENSG00000179172 | HNRNPCL1 | 92 | 50.357 |
| ENSG00000184672 | RALYL | 73 | 35.484 | ENSG00000124193 | SRSF6 | 53 | 32.877 |
| ENSG00000184672 | RALYL | 74 | 35.211 | ENSG00000147274 | RBMX | 90 | 35.211 |
| ENSG00000184672 | RALYL | 100 | 80.851 | ENSG00000125970 | RALY | 100 | 80.899 |
| ENSG00000184672 | RALYL | 60 | 43.137 | ENSG00000100650 | SRSF5 | 51 | 43.137 |
| ENSG00000184672 | RALYL | 92 | 80.769 | ENSG00000092199 | HNRNPC | 96 | 84.615 |
| Ensembl ID | Gene Symbol | Coverage | Identiy | Ortholog | Gene Symbol | Coverage | Identiy | Species |
|---|---|---|---|---|---|---|---|---|
| ENSG00000184672 | RALYL | 84 | 76.056 | ENSAPOG00000020641 | - | 84 | 76.056 | Acanthochromis_polyacanthus |
| ENSG00000184672 | RALYL | 100 | 91.489 | ENSAPOG00000005050 | RALYL | 83 | 90.588 | Acanthochromis_polyacanthus |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSAPOG00000006963 | hnrnpc | 85 | 52.015 | Acanthochromis_polyacanthus |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSACIG00000016568 | - | 99 | 50.373 | Amphilophus_citrinellus |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSACIG00000000270 | - | 95 | 50.523 | Amphilophus_citrinellus |
| ENSG00000184672 | RALYL | 93 | 88.608 | ENSACIG00000001335 | - | 67 | 87.500 | Amphilophus_citrinellus |
| ENSG00000184672 | RALYL | 93 | 75.949 | ENSAOCG00000021616 | - | 71 | 57.477 | Amphiprion_ocellaris |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSAOCG00000003896 | - | 70 | 57.727 | Amphiprion_ocellaris |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSAPEG00000001489 | - | 70 | 57.727 | Amphiprion_percula |
| ENSG00000184672 | RALYL | 91 | 88.312 | ENSAPEG00000021452 | - | 74 | 88.312 | Amphiprion_percula |
| ENSG00000184672 | RALYL | 100 | 84.706 | ENSAPEG00000010582 | - | 83 | 84.706 | Amphiprion_percula |
| ENSG00000184672 | RALYL | 93 | 75.949 | ENSAPEG00000019962 | - | 97 | 47.222 | Amphiprion_percula |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSATEG00000011734 | - | 74 | 58.296 | Anabas_testudineus |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSATEG00000018006 | - | 95 | 50.763 | Anabas_testudineus |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSAPLG00000015574 | RALYL | 100 | 92.759 | Anas_platyrhynchos |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSACAG00000028464 | RALYL | 100 | 91.126 | Anolis_carolinensis |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSANAG00000023383 | RALYL | 100 | 99.013 | Aotus_nancymaae |
| ENSG00000184672 | RALYL | 92 | 78.205 | ENSACLG00000005707 | - | 91 | 49.638 | Astatotilapia_calliptera |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSACLG00000013636 | - | 88 | 51.481 | Astatotilapia_calliptera |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSACLG00000000995 | - | 92 | 49.648 | Astatotilapia_calliptera |
| ENSG00000184672 | RALYL | 58 | 77.551 | ENSAMXG00000042946 | hnrnpc | 95 | 53.439 | Astyanax_mexicanus |
| ENSG00000184672 | RALYL | 99 | 73.810 | ENSAMXG00000034915 | zgc:55733 | 96 | 50.545 | Astyanax_mexicanus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSCJAG00000000402 | RALYL | 100 | 99.013 | Callithrix_jacchus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSCPOG00000012314 | RALYL | 100 | 95.548 | Cavia_porcellus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSCCAG00000025003 | RALYL | 100 | 99.013 | Cebus_capucinus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSCATG00000042145 | RALYL | 100 | 100.000 | Cercocebus_atys |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSCLAG00000005228 | RALYL | 100 | 96.233 | Chinchilla_lanigera |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSCHOG00000013228 | RALYL | 78 | 90.435 | Choloepus_hoffmanni |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSCANG00000039144 | RALYL | 100 | 99.671 | Colobus_angolensis_palliatus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSCGRG00001009241 | Ralyl | 100 | 95.563 | Cricetulus_griseus_chok1gshd |
| ENSG00000184672 | RALYL | 92 | 80.769 | ENSCSEG00000019406 | - | 75 | 80.769 | Cynoglossus_semilaevis |
| ENSG00000184672 | RALYL | 93 | 75.949 | ENSCSEG00000021349 | - | 89 | 48.763 | Cynoglossus_semilaevis |
| ENSG00000184672 | RALYL | 92 | 76.923 | ENSCSEG00000019769 | - | 71 | 56.744 | Cynoglossus_semilaevis |
| ENSG00000184672 | RALYL | 93 | 74.684 | ENSCVAG00000002797 | - | 90 | 45.926 | Cyprinodon_variegatus |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSCVAG00000016981 | - | 65 | 56.944 | Cyprinodon_variegatus |
| ENSG00000184672 | RALYL | 99 | 72.619 | ENSDARG00000103845 | zgc:55733 | 96 | 50.360 | Danio_rerio |
| ENSG00000184672 | RALYL | 92 | 74.359 | ENSDARG00000053810 | hnrnpc | 69 | 54.545 | Danio_rerio |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSDORG00000003721 | Ralyl | 100 | 95.563 | Dipodomys_ordii |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSETEG00000014028 | RALYL | 98 | 94.709 | Echinops_telfairi |
| ENSG00000184672 | RALYL | 92 | 73.077 | ENSEBUG00000012363 | hnrnpc | 69 | 47.755 | Eptatretus_burgeri |
| ENSG00000184672 | RALYL | 100 | 72.941 | ENSEBUG00000006049 | RALYL | 94 | 49.632 | Eptatretus_burgeri |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSEEUG00000007202 | RALYL | 100 | 94.792 | Erinaceus_europaeus |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSELUG00000007714 | - | 96 | 48.727 | Esox_lucius |
| ENSG00000184672 | RALYL | 99 | 71.429 | ENSELUG00000004132 | - | 74 | 55.804 | Esox_lucius |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSFALG00000008647 | RALYL | 100 | 92.130 | Ficedula_albicollis |
| ENSG00000184672 | RALYL | 91 | 87.013 | ENSFHEG00000000223 | - | 79 | 87.013 | Fundulus_heteroclitus |
| ENSG00000184672 | RALYL | 91 | 76.623 | ENSFHEG00000003605 | - | 87 | 50.185 | Fundulus_heteroclitus |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSFHEG00000007278 | - | 69 | 53.478 | Fundulus_heteroclitus |
| ENSG00000184672 | RALYL | 100 | 90.588 | ENSFHEG00000017466 | - | 91 | 67.442 | Fundulus_heteroclitus |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSGALG00000036938 | RALYL | 100 | 91.864 | Gallus_gallus |
| ENSG00000184672 | RALYL | 100 | 89.412 | ENSGAFG00000018541 | RALYL | 100 | 77.670 | Gambusia_affinis |
| ENSG00000184672 | RALYL | 94 | 87.500 | ENSGAFG00000020052 | - | 89 | 79.348 | Gambusia_affinis |
| ENSG00000184672 | RALYL | 91 | 76.623 | ENSGAFG00000020486 | - | 98 | 48.352 | Gambusia_affinis |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSGAFG00000020992 | - | 99 | 48.727 | Gambusia_affinis |
| ENSG00000184672 | RALYL | 98 | 71.739 | ENSGACG00000017772 | - | 82 | 56.744 | Gasterosteus_aculeatus |
| ENSG00000184672 | RALYL | 93 | 72.152 | ENSGACG00000018932 | - | 84 | 50.228 | Gasterosteus_aculeatus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSGGOG00000000841 | RALYL | 100 | 100.000 | Gorilla_gorilla |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSHBUG00000018342 | - | 92 | 50.000 | Haplochromis_burtoni |
| ENSG00000184672 | RALYL | 92 | 79.487 | ENSHBUG00000010527 | - | 87 | 47.000 | Haplochromis_burtoni |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSHGLG00000004866 | RALYL | 100 | 95.548 | Heterocephalus_glaber_female |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSHGLG00100017480 | RALYL | 100 | 95.548 | Heterocephalus_glaber_male |
| ENSG00000184672 | RALYL | 100 | 85.882 | ENSHCOG00000002851 | - | 77 | 85.882 | Hippocampus_comes |
| ENSG00000184672 | RALYL | 92 | 76.923 | ENSHCOG00000012333 | hnrnpc | 76 | 54.468 | Hippocampus_comes |
| ENSG00000184672 | RALYL | 92 | 88.462 | ENSHCOG00000012719 | - | 83 | 81.818 | Hippocampus_comes |
| ENSG00000184672 | RALYL | 92 | 76.923 | ENSIPUG00000011271 | hnrnpc | 95 | 54.260 | Ictalurus_punctatus |
| ENSG00000184672 | RALYL | 99 | 72.619 | ENSIPUG00000008666 | zgc:55733 | 92 | 60.476 | Ictalurus_punctatus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSSTOG00000012583 | RALYL | 100 | 95.833 | Ictidomys_tridecemlineatus |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSKMAG00000011170 | - | 83 | 51.866 | Kryptolebias_marmoratus |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSKMAG00000018265 | - | 66 | 58.605 | Kryptolebias_marmoratus |
| ENSG00000184672 | RALYL | 92 | 75.641 | ENSLBEG00000006193 | - | 84 | 49.638 | Labrus_bergylta |
| ENSG00000184672 | RALYL | 93 | 75.949 | ENSLBEG00000024418 | - | 74 | 53.070 | Labrus_bergylta |
| ENSG00000184672 | RALYL | 91 | 76.623 | ENSLBEG00000006979 | - | 73 | 55.760 | Labrus_bergylta |
| ENSG00000184672 | RALYL | 100 | 94.118 | ENSLACG00000000135 | - | 100 | 91.011 | Latimeria_chalumnae |
| ENSG00000184672 | RALYL | 99 | 71.429 | ENSLOCG00000002560 | hnrnpc | 84 | 49.393 | Lepisosteus_oculatus |
| ENSG00000184672 | RALYL | 92 | 80.769 | ENSMMUG00000003920 | - | 97 | 75.532 | Macaca_mulatta |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSMNEG00000033288 | RALYL | 100 | 100.000 | Macaca_nemestrina |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSMLEG00000038284 | RALYL | 100 | 100.000 | Mandrillus_leucophaeus |
| ENSG00000184672 | RALYL | 100 | 90.588 | ENSMAMG00000011137 | RALYL | 99 | 81.818 | Mastacembelus_armatus |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSMAMG00000008033 | - | 74 | 58.411 | Mastacembelus_armatus |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSMAMG00000012236 | - | 74 | 58.716 | Mastacembelus_armatus |
| ENSG00000184672 | RALYL | 92 | 78.205 | ENSMZEG00005024249 | - | 85 | 49.638 | Maylandia_zebra |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSMZEG00005020460 | - | 92 | 49.648 | Maylandia_zebra |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSMAUG00000015149 | Ralyl | 100 | 95.563 | Mesocricetus_auratus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSMICG00000005558 | RALYL | 100 | 97.153 | Microcebus_murinus |
| ENSG00000184672 | RALYL | 100 | 98.824 | ENSMOCG00000017371 | Ralyl | 100 | 94.198 | Microtus_ochrogaster |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSMMOG00000021465 | - | 95 | 51.254 | Mola_mola |
| ENSG00000184672 | RALYL | 96 | 84.146 | ENSMMOG00000004599 | - | 76 | 84.146 | Mola_mola |
| ENSG00000184672 | RALYL | 98 | 74.699 | ENSMMOG00000019520 | - | 99 | 52.809 | Mola_mola |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSMODG00000029239 | - | 89 | 93.478 | Monodelphis_domestica |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSMALG00000007520 | - | 95 | 50.545 | Monopterus_albus |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSMALG00000014623 | - | 85 | 49.291 | Monopterus_albus |
| ENSG00000184672 | RALYL | 100 | 98.824 | ENSMUSG00000039717 | Ralyl | 100 | 98.824 | Mus_musculus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSMPUG00000010724 | RALYL | 100 | 95.904 | Mustela_putorius_furo |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSNGAG00000013375 | Ralyl | 100 | 95.563 | Nannospalax_galili |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSNBRG00000004912 | - | 94 | 53.648 | Neolamprologus_brichardi |
| ENSG00000184672 | RALYL | 92 | 79.487 | ENSNBRG00000015676 | - | 87 | 46.667 | Neolamprologus_brichardi |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSNLEG00000002702 | RALYL | 96 | 100.000 | Nomascus_leucogenys |
| ENSG00000184672 | RALYL | 86 | 81.119 | ENSOPRG00000013650 | - | 100 | 87.958 | Ochotona_princeps |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSODEG00000001012 | RALYL | 100 | 95.205 | Octodon_degus |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSONIG00000007859 | - | 85 | 52.222 | Oreochromis_niloticus |
| ENSG00000184672 | RALYL | 92 | 79.487 | ENSONIG00000018350 | - | 69 | 56.757 | Oreochromis_niloticus |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSOANG00000030197 | - | 61 | 93.478 | Ornithorhynchus_anatinus |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSORLG00000009999 | - | 72 | 53.304 | Oryzias_latipes |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSORLG00000003144 | - | 66 | 56.306 | Oryzias_latipes |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSORLG00020018228 | hnrnpc | 72 | 53.304 | Oryzias_latipes_hni |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSORLG00015009460 | - | 66 | 56.306 | Oryzias_latipes_hsok |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSORLG00015019755 | - | 72 | 53.304 | Oryzias_latipes_hsok |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSOMEG00000022665 | - | 66 | 58.140 | Oryzias_melastigma |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSOMEG00000020488 | - | 72 | 54.185 | Oryzias_melastigma |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSPPAG00000038705 | RALYL | 100 | 100.000 | Pan_paniscus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSPPRG00000018612 | RALYL | 100 | 96.099 | Panthera_pardus |
| ENSG00000184672 | RALYL | 100 | 97.872 | ENSPTIG00000020331 | RALYL | 100 | 95.745 | Panthera_tigris_altaica |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSPTRG00000020381 | RALYL | 100 | 100.000 | Pan_troglodytes |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSPANG00000029305 | RALYL | 100 | 100.000 | Papio_anubis |
| ENSG00000184672 | RALYL | 99 | 73.810 | ENSPKIG00000008592 | - | 86 | 50.355 | Paramormyrops_kingsleyae |
| ENSG00000184672 | RALYL | 100 | 90.588 | ENSPKIG00000023807 | RALYL | 87 | 90.588 | Paramormyrops_kingsleyae |
| ENSG00000184672 | RALYL | 92 | 79.487 | ENSPKIG00000022837 | - | 85 | 52.788 | Paramormyrops_kingsleyae |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSPSIG00000011135 | RALYL | 100 | 90.847 | Pelodiscus_sinensis |
| ENSG00000184672 | RALYL | 91 | 81.818 | ENSPMGG00000017774 | - | 88 | 50.935 | Periophthalmus_magnuspinnatus |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSPMGG00000013144 | - | 98 | 50.376 | Periophthalmus_magnuspinnatus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSPEMG00000016447 | Ralyl | 100 | 95.222 | Peromyscus_maniculatus_bairdii |
| ENSG00000184672 | RALYL | 100 | 87.234 | ENSPMAG00000002407 | RALYL | 100 | 69.432 | Petromyzon_marinus |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSPCIG00000023938 | - | 80 | 93.478 | Phascolarctos_cinereus |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSPFOG00000001583 | - | 73 | 54.626 | Poecilia_formosa |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSPFOG00000002643 | - | 66 | 57.277 | Poecilia_formosa |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSPLAG00000000061 | - | 65 | 57.277 | Poecilia_latipinna |
| ENSG00000184672 | RALYL | 91 | 76.623 | ENSPLAG00000015305 | - | 73 | 53.982 | Poecilia_latipinna |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSPMEG00000004802 | - | 73 | 54.425 | Poecilia_mexicana |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSPMEG00000016524 | - | 66 | 57.277 | Poecilia_mexicana |
| ENSG00000184672 | RALYL | 91 | 79.221 | ENSPREG00000007928 | - | 73 | 54.867 | Poecilia_reticulata |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSPREG00000001852 | - | 71 | 56.808 | Poecilia_reticulata |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSPPYG00000018710 | - | 100 | 100.000 | Pongo_abelii |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSPVAG00000010821 | RALYL | 99 | 89.041 | Pteropus_vampyrus |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSPNYG00000020293 | - | 89 | 52.697 | Pundamilia_nyererei |
| ENSG00000184672 | RALYL | 92 | 79.487 | ENSPNYG00000003747 | - | 93 | 47.000 | Pundamilia_nyererei |
| ENSG00000184672 | RALYL | 92 | 76.923 | ENSPNAG00000019200 | zgc:55733 | 93 | 49.826 | Pygocentrus_nattereri |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSPNAG00000015686 | hnrnpc | 74 | 55.909 | Pygocentrus_nattereri |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSRROG00000023925 | RALYL | 100 | 100.000 | Rhinopithecus_roxellana |
| ENSG00000184672 | RALYL | 92 | 79.487 | ENSSFOG00015012924 | - | 96 | 67.368 | Scleropages_formosus |
| ENSG00000184672 | RALYL | 92 | 76.923 | ENSSFOG00015012015 | - | 99 | 50.558 | Scleropages_formosus |
| ENSG00000184672 | RALYL | 91 | 75.325 | ENSSMAG00000013592 | - | 67 | 57.277 | Scophthalmus_maximus |
| ENSG00000184672 | RALYL | 99 | 72.619 | ENSSMAG00000011147 | - | 76 | 58.768 | Scophthalmus_maximus |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSSDUG00000001723 | - | 87 | 44.737 | Seriola_dumerili |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSSDUG00000006730 | - | 86 | 48.110 | Seriola_dumerili |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSSLDG00000008069 | - | 75 | 77.215 | Seriola_lalandi_dorsalis |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSSLDG00000007454 | - | 68 | 59.242 | Seriola_lalandi_dorsalis |
| ENSG00000184672 | RALYL | 93 | 77.215 | ENSSLDG00000007474 | - | 91 | 50.545 | Seriola_lalandi_dorsalis |
| ENSG00000184672 | RALYL | 100 | 97.872 | ENSSARG00000002374 | RALYL | 100 | 73.776 | Sorex_araneus |
| ENSG00000184672 | RALYL | 93 | 75.949 | ENSSPAG00000013996 | - | 67 | 57.944 | Stegastes_partitus |
| ENSG00000184672 | RALYL | 100 | 84.706 | ENSSPAG00000014679 | - | 89 | 84.706 | Stegastes_partitus |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSSPAG00000023204 | - | 72 | 56.889 | Stegastes_partitus |
| ENSG00000184672 | RALYL | 100 | 96.471 | ENSTGUG00000011689 | RALYL | 100 | 91.724 | Taeniopygia_guttata |
| ENSG00000184672 | RALYL | 93 | 74.684 | ENSTRUG00000020158 | - | 94 | 47.038 | Takifugu_rubripes |
| ENSG00000184672 | RALYL | 93 | 78.481 | ENSTRUG00000002153 | - | 78 | 73.333 | Takifugu_rubripes |
| ENSG00000184672 | RALYL | 100 | 89.412 | ENSTRUG00000021320 | RALYL | 92 | 89.412 | Takifugu_rubripes |
| ENSG00000184672 | RALYL | 93 | 79.747 | ENSTNIG00000000334 | - | 99 | 51.103 | Tetraodon_nigroviridis |
| ENSG00000184672 | RALYL | 93 | 74.684 | ENSTNIG00000007427 | - | 73 | 51.786 | Tetraodon_nigroviridis |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSTBEG00000005551 | RALYL | 100 | 85.771 | Tupaia_belangeri |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSTTRG00000015762 | RALYL | 100 | 91.468 | Tursiops_truncatus |
| ENSG00000184672 | RALYL | 99 | 76.190 | ENSUAMG00000006020 | - | 88 | 76.190 | Ursus_americanus |
| ENSG00000184672 | RALYL | 100 | 100.000 | ENSVPAG00000008501 | RALYL | 100 | 91.034 | Vicugna_pacos |
| ENSG00000184672 | RALYL | 92 | 78.205 | ENSXCOG00000006552 | - | 88 | 46.403 | Xiphophorus_couchianus |
| ENSG00000184672 | RALYL | 93 | 75.949 | ENSXCOG00000019434 | - | 78 | 55.191 | Xiphophorus_couchianus |
| ENSG00000184672 | RALYL | 91 | 87.013 | ENSXCOG00000016096 | - | 71 | 82.927 | Xiphophorus_couchianus |
| ENSG00000184672 | RALYL | 91 | 76.623 | ENSXMAG00000022849 | - | 69 | 56.338 | Xiphophorus_maculatus |
| ENSG00000184672 | RALYL | 91 | 77.922 | ENSXMAG00000009333 | - | 73 | 53.540 | Xiphophorus_maculatus |
| Go ID | Go_term | PubmedID | Evidence | Category |
|---|---|---|---|---|
| GO:0003723 | RNA binding | 22658674. | HDA | Function |
| GO:0003723 | RNA binding | 21873635. | IBA | Function |
| GO:0005515 | protein binding | 16189514.19447967.25416956. | IPI | Function |
| GO:0005634 | nucleus | 21873635. | IBA | Component |
| GO:0042802 | identical protein binding | 16189514.19447967.25416956. | IPI | Function |