
| Cancer | Chr | Position | Mutation Type | dbSNP | Protein-change | Allele Freq | RBD |
|---|---|---|---|---|---|---|---|
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| BRCA | |||||||
| CESC | |||||||
| CESC | |||||||
| CESC | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| ESCA | |||||||
| ESCA | |||||||
| GBM | |||||||
| HNSC | |||||||
| HNSC | |||||||
| KICH | |||||||
| KIRC | |||||||
| KIRP | |||||||
| KIRP | |||||||
| KIRP | |||||||
| LGG | |||||||
| LIHC | |||||||
| LIHC | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| LUSC | |||||||
| OV | |||||||
| PAAD | |||||||
| PAAD | |||||||
| PAAD | |||||||
| PAAD | |||||||
| PRAD | |||||||
| SARC | |||||||
| SARC | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| STAD | |||||||
| THCA | |||||||
| THCA | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCS |
| Cancer | Type | Freq | Q-value |
|---|---|---|---|
| LUSC | |||
| LUSC | |||
| TGCT | |||
| THCA |
| Cancer | P-value | Q-value |
|---|---|---|
| KIRC | ||
| STAD | ||
| MESO | ||
| ACC | ||
| SKCM | ||
| COAD | ||
| PAAD | ||
| BLCA | ||
| READ | ||
| UCEC | ||
| LIHC | ||
| LGG | ||
| UVM |
| Protein ID | Domain | Pfam ID | E-value | Domain number | Total number |
|---|---|---|---|---|---|
| ENSP00000378541 | MMR_HSR1 | PF01926.23 | 4.2e-06 | 1 | 1 |
| ENSP00000414488 | MMR_HSR1 | PF01926.23 | 5.3e-06 | 1 | 1 |
| ENSP00000391311 | MMR_HSR1 | PF01926.23 | 6.9e-06 | 1 | 1 |
| ENSP00000384535 | MMR_HSR1 | PF01926.23 | 7e-06 | 1 | 1 |
| ENSP00000372515 | MMR_HSR1 | PF01926.23 | 7.3e-06 | 1 | 1 |
| ENSP00000394541 | MMR_HSR1 | PF01926.23 | 7.4e-06 | 1 | 1 |
| ENSP00000408678 | MMR_HSR1 | PF01926.23 | 9.7e-06 | 1 | 1 |
| Transcript ID | Name | Length | RefSeq ID | Protein ID | Length | RefSeq ID | UniportKB ID |
|---|---|---|---|---|---|---|---|
| ENST00000431124 | SEPT5-207 | 920 | - | ENSP00000414488 | 304 (aa) | - | E7EPG2 |
| ENST00000438754 | SEPT5-208 | 2241 | - | ENSP00000394541 | 346 (aa) | - | Q99719 |
| ENST00000406172 | SEPT5-203 | 1996 | - | ENSP00000385356 | 248 (aa) | - | E7EQM7 |
| ENST00000383045 | SEPT5-201 | 1588 | - | ENSP00000372515 | 378 (aa) | - | G3XAH0 |
| ENST00000480423 | SEPT5-212 | 563 | - | - | - (aa) | - | - |
| ENST00000490204 | SEPT5-213 | 774 | - | - | - (aa) | - | - |
| ENST00000455784 | SEPT5-209 | 2070 | - | ENSP00000391311 | 369 (aa) | - | Q99719 |
| ENST00000406395 | SEPT5-204 | 1974 | - | ENSP00000384535 | 337 (aa) | - | E7EX32 |
| ENST00000470814 | SEPT5-210 | 2353 | - | - | - (aa) | - | - |
| ENST00000412544 | SEPT5-205 | 1108 | - | ENSP00000408678 | 301 (aa) | - | C9JM82 |
| ENST00000395109 | SEPT5-202 | 794 | - | ENSP00000378541 | 180 (aa) | - | F8W9E5 |
| ENST00000477238 | SEPT5-211 | 577 | - | - | - (aa) | - | - |
| ENST00000413258 | SEPT5-206 | 318 | - | ENSP00000404673 | 106 (aa) | - | H7C299 |
| Pathway ID | Pathway Name | Source |
|---|---|---|
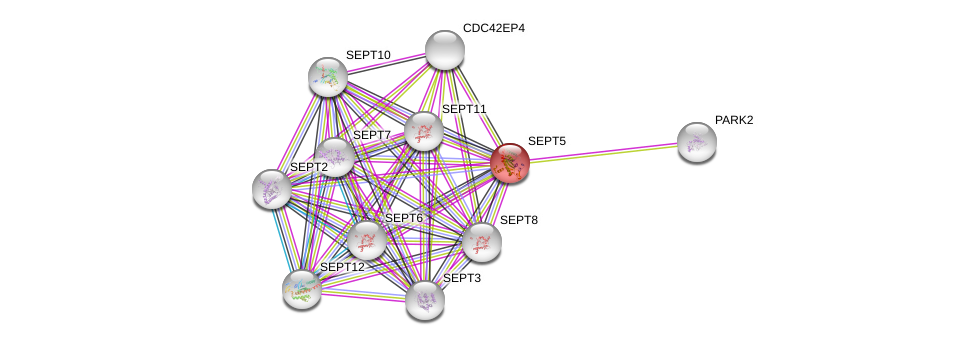
| hsa05012 | Parkinson disease | KEGG |
| Ensembl ID | Gene Symbol | Coverage | Identiy | Ortholog | Gene Symbol | Coverage | Identiy | Species |
|---|---|---|---|---|---|---|---|---|
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSAPOG00000021568 | sept5a | 99 | 85.366 | Acanthochromis_polyacanthus |
| ENSG00000184702 | SEPT5 | 100 | 87.222 | ENSAPOG00000004707 | sept5b | 98 | 79.133 | Acanthochromis_polyacanthus |
| ENSG00000184702 | SEPT5 | 100 | 98.007 | ENSAMEG00000011509 | - | 87 | 94.215 | Ailuropoda_melanoleuca |
| ENSG00000184702 | SEPT5 | 100 | 87.778 | ENSACIG00000000964 | - | 99 | 78.693 | Amphilophus_citrinellus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSACIG00000006554 | sept5a | 97 | 89.143 | Amphilophus_citrinellus |
| ENSG00000184702 | SEPT5 | 100 | 87.222 | ENSAOCG00000019424 | sept5b | 93 | 85.387 | Amphiprion_ocellaris |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSAOCG00000015062 | sept5a | 99 | 85.366 | Amphiprion_ocellaris |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSAPEG00000017611 | sept5a | 99 | 85.366 | Amphiprion_percula |
| ENSG00000184702 | SEPT5 | 100 | 87.222 | ENSAPEG00000008250 | sept5b | 98 | 79.675 | Amphiprion_percula |
| ENSG00000184702 | SEPT5 | 100 | 86.667 | ENSATEG00000006356 | sept5b | 98 | 83.056 | Anabas_testudineus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSATEG00000020583 | sept5a | 99 | 85.637 | Anabas_testudineus |
| ENSG00000184702 | SEPT5 | 100 | 95.017 | ENSAPLG00000016224 | - | 100 | 94.587 | Anas_platyrhynchos |
| ENSG00000184702 | SEPT5 | 100 | 95.681 | ENSACAG00000000290 | - | 82 | 93.767 | Anolis_carolinensis |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSANAG00000025950 | - | 100 | 100.000 | Aotus_nancymaae |
| ENSG00000184702 | SEPT5 | 100 | 86.667 | ENSACLG00000008299 | sept5b | 99 | 80.759 | Astatotilapia_calliptera |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSACLG00000011272 | sept5a | 97 | 89.112 | Astatotilapia_calliptera |
| ENSG00000184702 | SEPT5 | 100 | 88.889 | ENSAMXG00000006159 | sept5a | 99 | 85.366 | Astyanax_mexicanus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSBTAG00000008320 | SEPT5 | 100 | 98.413 | Bos_taurus |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSCJAG00000031147 | - | 100 | 100.000 | Callithrix_jacchus |
| ENSG00000184702 | SEPT5 | 100 | 99.336 | ENSCAFG00000014533 | SEPT5 | 100 | 99.206 | Canis_familiaris |
| ENSG00000184702 | SEPT5 | 100 | 99.336 | ENSCAFG00020007897 | - | 100 | 99.206 | Canis_lupus_dingo |
| ENSG00000184702 | SEPT5 | 100 | 95.580 | ENSCHIG00000000129 | - | 92 | 91.185 | Capra_hircus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSTSYG00000037551 | - | 100 | 98.677 | Carlito_syrichta |
| ENSG00000184702 | SEPT5 | 100 | 97.778 | ENSCAPG00000006545 | - | 100 | 98.697 | Cavia_aperea |
| ENSG00000184702 | SEPT5 | 100 | 98.677 | ENSCPOG00000010483 | - | 100 | 98.677 | Cavia_porcellus |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSCCAG00000036918 | - | 100 | 100.000 | Cebus_capucinus |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSCATG00000040593 | - | 100 | 100.000 | Cercocebus_atys |
| ENSG00000184702 | SEPT5 | 100 | 98.148 | ENSCLAG00000015333 | - | 100 | 98.148 | Chinchilla_lanigera |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSCSAG00000011732 | - | 100 | 100.000 | Chlorocebus_sabaeus |
| ENSG00000184702 | SEPT5 | 100 | 96.111 | ENSCPBG00000010660 | - | 100 | 94.851 | Chrysemys_picta_bellii |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSCANG00000030373 | - | 100 | 99.464 | Colobus_angolensis_palliatus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSCGRG00001018409 | - | 100 | 98.916 | Cricetulus_griseus_chok1gshd |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSCGRG00000011656 | - | 93 | 96.000 | Cricetulus_griseus_crigri |
| ENSG00000184702 | SEPT5 | 100 | 91.111 | ENSCSEG00000019641 | sept5a | 99 | 85.100 | Cynoglossus_semilaevis |
| ENSG00000184702 | SEPT5 | 100 | 89.444 | ENSCSEG00000009877 | - | 91 | 87.162 | Cynoglossus_semilaevis |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSCVAG00000017986 | sept5a | 98 | 87.115 | Cyprinodon_variegatus |
| ENSG00000184702 | SEPT5 | 100 | 87.778 | ENSDARG00000013843 | sept5a | 90 | 84.282 | Danio_rerio |
| ENSG00000184702 | SEPT5 | 100 | 87.778 | ENSDARG00000036031 | sept5b | 100 | 84.133 | Danio_rerio |
| ENSG00000184702 | SEPT5 | 100 | 99.336 | ENSDORG00000013930 | - | 100 | 99.187 | Dipodomys_ordii |
| ENSG00000184702 | SEPT5 | 100 | 98.007 | ENSEASG00005010502 | - | 96 | 97.826 | Equus_asinus_asinus |
| ENSG00000184702 | SEPT5 | 100 | 98.671 | ENSECAG00000020718 | - | 70 | 98.374 | Equus_caballus |
| ENSG00000184702 | SEPT5 | 100 | 89.444 | ENSELUG00000009028 | sept5a | 99 | 84.085 | Esox_lucius |
| ENSG00000184702 | SEPT5 | 100 | 87.778 | ENSELUG00000005164 | sept5b | 99 | 82.289 | Esox_lucius |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSFCAG00000035936 | - | 100 | 97.561 | Felis_catus |
| ENSG00000184702 | SEPT5 | 100 | 96.111 | ENSFALG00000008028 | - | 100 | 93.767 | Ficedula_albicollis |
| ENSG00000184702 | SEPT5 | 100 | 98.671 | ENSFDAG00000014909 | - | 100 | 98.588 | Fukomys_damarensis |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSFHEG00000018533 | sept5a | 98 | 88.286 | Fundulus_heteroclitus |
| ENSG00000184702 | SEPT5 | 100 | 72.848 | ENSFHEG00000003693 | sept5b | 99 | 74.013 | Fundulus_heteroclitus |
| ENSG00000184702 | SEPT5 | 100 | 95.556 | ENSGALG00000001688 | SEPT5 | 100 | 92.954 | Gallus_gallus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSGAFG00000000767 | sept5a | 98 | 89.398 | Gambusia_affinis |
| ENSG00000184702 | SEPT5 | 100 | 85.556 | ENSGAFG00000007663 | sept5b | 90 | 79.545 | Gambusia_affinis |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSGACG00000005538 | sept5a | 98 | 88.761 | Gasterosteus_aculeatus |
| ENSG00000184702 | SEPT5 | 100 | 96.111 | ENSGAGG00000014385 | - | 100 | 92.670 | Gopherus_agassizii |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSGGOG00000016545 | - | 100 | 100.000 | Gorilla_gorilla |
| ENSG00000184702 | SEPT5 | 100 | 87.222 | ENSHBUG00000007908 | sept5b | 99 | 81.030 | Haplochromis_burtoni |
| ENSG00000184702 | SEPT5 | 68 | 91.803 | ENSHBUG00000013963 | sept5a | 93 | 92.617 | Haplochromis_burtoni |
| ENSG00000184702 | SEPT5 | 100 | 98.671 | ENSHGLG00000013741 | - | 100 | 98.645 | Heterocephalus_glaber_female |
| ENSG00000184702 | SEPT5 | 100 | 91.111 | ENSHCOG00000005881 | sept5a | 99 | 85.366 | Hippocampus_comes |
| ENSG00000184702 | SEPT5 | 100 | 78.405 | ENSIPUG00000025006 | sept5a | 94 | 59.593 | Ictalurus_punctatus |
| ENSG00000184702 | SEPT5 | 100 | 99.668 | ENSSTOG00000010725 | - | 100 | 99.471 | Ictidomys_tridecemlineatus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSJJAG00000000091 | - | 100 | 98.916 | Jaculus_jaculus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSKMAG00000003408 | sept5a | 99 | 85.908 | Kryptolebias_marmoratus |
| ENSG00000184702 | SEPT5 | 100 | 90.556 | ENSLBEG00000025389 | sept5a | 99 | 85.366 | Labrus_bergylta |
| ENSG00000184702 | SEPT5 | 100 | 85.000 | ENSLBEG00000028216 | - | 92 | 83.815 | Labrus_bergylta |
| ENSG00000184702 | SEPT5 | 100 | 92.222 | ENSLACG00000015559 | sept5b | 100 | 84.304 | Latimeria_chalumnae |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSLAFG00000028004 | - | 100 | 98.113 | Loxodonta_africana |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSMFAG00000038117 | - | 100 | 100.000 | Macaca_fascicularis |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSMMUG00000017134 | - | 100 | 100.000 | Macaca_mulatta |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSMNEG00000036305 | - | 100 | 100.000 | Macaca_nemestrina |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSMLEG00000029088 | - | 100 | 100.000 | Mandrillus_leucophaeus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSMAMG00000014426 | sept5a | 99 | 85.637 | Mastacembelus_armatus |
| ENSG00000184702 | SEPT5 | 100 | 86.667 | ENSMAMG00000006704 | sept5b | 99 | 82.102 | Mastacembelus_armatus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSMZEG00005009762 | sept5a | 97 | 87.675 | Maylandia_zebra |
| ENSG00000184702 | SEPT5 | 100 | 86.667 | ENSMZEG00005011980 | sept5b | 96 | 83.862 | Maylandia_zebra |
| ENSG00000184702 | SEPT5 | 100 | 93.069 | ENSMGAG00000001319 | - | 96 | 92.938 | Meleagris_gallopavo |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSMAUG00000014786 | - | 100 | 98.916 | Mesocricetus_auratus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSMICG00000028407 | - | 100 | 98.677 | Microcebus_murinus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSMOCG00000000071 | - | 100 | 98.916 | Microtus_ochrogaster |
| ENSG00000184702 | SEPT5 | 100 | 84.444 | ENSMMOG00000010567 | sept5b | 99 | 78.049 | Mola_mola |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSMMOG00000011704 | sept5a | 99 | 89.112 | Mola_mola |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSMALG00000020359 | sept5a | 98 | 93.836 | Monopterus_albus |
| ENSG00000184702 | SEPT5 | 100 | 84.444 | ENSMALG00000005830 | sept5b | 99 | 82.633 | Monopterus_albus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | MGP_CAROLIEiJ_G0020551 | - | 100 | 98.916 | Mus_caroli |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSMUSG00000072214 | Sept5 | 100 | 98.942 | Mus_musculus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSMUSG00000116594 | AC133488.1 | 99 | 99.167 | Mus_musculus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | MGP_PahariEiJ_G0016256 | - | 100 | 98.916 | Mus_pahari |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | MGP_SPRETEiJ_G0021446 | - | 100 | 98.942 | Mus_spretus |
| ENSG00000184702 | SEPT5 | 100 | 99.336 | ENSMPUG00000003893 | - | 100 | 99.068 | Mustela_putorius_furo |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSMLUG00000002506 | - | 96 | 98.942 | Myotis_lucifugus |
| ENSG00000184702 | SEPT5 | 100 | 99.336 | ENSNGAG00000013782 | - | 100 | 99.187 | Nannospalax_galili |
| ENSG00000184702 | SEPT5 | 100 | 86.667 | ENSNBRG00000004761 | sept5b | 96 | 83.862 | Neolamprologus_brichardi |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSNBRG00000018717 | sept5a | 97 | 89.112 | Neolamprologus_brichardi |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSNLEG00000005511 | - | 100 | 99.074 | Nomascus_leucogenys |
| ENSG00000184702 | SEPT5 | 100 | 96.111 | ENSMEUG00000015556 | - | 100 | 95.726 | Notamacropus_eugenii |
| ENSG00000184702 | SEPT5 | 100 | 95.349 | ENSOPRG00000007708 | - | 100 | 95.726 | Ochotona_princeps |
| ENSG00000184702 | SEPT5 | 100 | 98.671 | ENSODEG00000016498 | - | 100 | 98.645 | Octodon_degus |
| ENSG00000184702 | SEPT5 | 100 | 87.222 | ENSONIG00000011388 | sept5b | 99 | 81.301 | Oreochromis_niloticus |
| ENSG00000184702 | SEPT5 | 100 | 83.889 | ENSORLG00000001081 | sept5b | 98 | 81.440 | Oryzias_latipes |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSORLG00000024104 | sept5a | 98 | 89.112 | Oryzias_latipes |
| ENSG00000184702 | SEPT5 | 100 | 83.333 | ENSORLG00020016872 | sept5b | 94 | 82.808 | Oryzias_latipes_hni |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSORLG00020005445 | sept5a | 98 | 89.112 | Oryzias_latipes_hni |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSORLG00015015192 | sept5a | 99 | 85.637 | Oryzias_latipes_hsok |
| ENSG00000184702 | SEPT5 | 100 | 83.333 | ENSORLG00015003711 | sept5b | 98 | 81.440 | Oryzias_latipes_hsok |
| ENSG00000184702 | SEPT5 | 99 | 84.270 | ENSOMEG00000022819 | sept5b | 89 | 82.963 | Oryzias_melastigma |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSOMEG00000017874 | sept5a | 99 | 89.112 | Oryzias_melastigma |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSOGAG00000026834 | - | 99 | 95.640 | Otolemur_garnettii |
| ENSG00000184702 | SEPT5 | 66 | 95.798 | ENSOARG00000001444 | - | 90 | 95.798 | Ovis_aries |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSPPAG00000042516 | - | 100 | 100.000 | Pan_paniscus |
| ENSG00000184702 | SEPT5 | 100 | 99.668 | ENSPPRG00000002930 | - | 100 | 99.471 | Panthera_pardus |
| ENSG00000184702 | SEPT5 | 100 | 99.668 | ENSPTIG00000019967 | - | 100 | 99.471 | Panthera_tigris_altaica |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSPTRG00000014066 | - | 100 | 100.000 | Pan_troglodytes |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSPANG00000001023 | - | 100 | 100.000 | Papio_anubis |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSPKIG00000024641 | sept5b | 99 | 87.805 | Paramormyrops_kingsleyae |
| ENSG00000184702 | SEPT5 | 100 | 95.681 | ENSPSIG00000002889 | - | 100 | 95.342 | Pelodiscus_sinensis |
| ENSG00000184702 | SEPT5 | 100 | 87.222 | ENSPMGG00000021306 | sept5b | 98 | 83.333 | Periophthalmus_magnuspinnatus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSPMGG00000018375 | sept5a | 99 | 86.339 | Periophthalmus_magnuspinnatus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSPEMG00000013636 | - | 100 | 98.870 | Peromyscus_maniculatus_bairdii |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSPFOG00000011811 | sept5a | 99 | 86.179 | Poecilia_formosa |
| ENSG00000184702 | SEPT5 | 100 | 85.556 | ENSPFOG00000016894 | sept5b | 98 | 81.111 | Poecilia_formosa |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSPLAG00000018547 | sept5a | 98 | 89.685 | Poecilia_latipinna |
| ENSG00000184702 | SEPT5 | 100 | 85.000 | ENSPLAG00000005877 | sept5b | 99 | 79.620 | Poecilia_latipinna |
| ENSG00000184702 | SEPT5 | 100 | 85.556 | ENSPMEG00000017826 | sept5b | 96 | 79.184 | Poecilia_mexicana |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSPMEG00000004070 | sept5a | 99 | 85.908 | Poecilia_mexicana |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSPREG00000011585 | sept5a | 99 | 85.450 | Poecilia_reticulata |
| ENSG00000184702 | SEPT5 | 100 | 85.083 | ENSPREG00000013868 | sept5b | 97 | 79.245 | Poecilia_reticulata |
| ENSG00000184702 | SEPT5 | 100 | 99.444 | ENSPPYG00000011583 | - | 87 | 99.550 | Pongo_abelii |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSPCOG00000012479 | - | 100 | 98.916 | Propithecus_coquereli |
| ENSG00000184702 | SEPT5 | 100 | 99.336 | ENSPVAG00000012974 | - | 93 | 93.634 | Pteropus_vampyrus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSPNYG00000008373 | sept5a | 97 | 89.112 | Pundamilia_nyererei |
| ENSG00000184702 | SEPT5 | 100 | 86.667 | ENSPNYG00000010341 | sept5b | 99 | 83.621 | Pundamilia_nyererei |
| ENSG00000184702 | SEPT5 | 100 | 89.369 | ENSPNAG00000001270 | sept5b | 99 | 86.179 | Pygocentrus_nattereri |
| ENSG00000184702 | SEPT5 | 100 | 89.444 | ENSPNAG00000007785 | sept5a | 99 | 85.908 | Pygocentrus_nattereri |
| ENSG00000184702 | SEPT5 | 100 | 98.677 | ENSRNOG00000029912 | Sept5 | 100 | 98.677 | Rattus_norvegicus |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSRBIG00000041362 | - | 100 | 100.000 | Rhinopithecus_bieti |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSRROG00000041246 | - | 100 | 100.000 | Rhinopithecus_roxellana |
| ENSG00000184702 | SEPT5 | 100 | 100.000 | ENSSBOG00000027910 | - | 100 | 100.000 | Saimiri_boliviensis_boliviensis |
| ENSG00000184702 | SEPT5 | 100 | 96.111 | ENSSHAG00000011753 | - | 92 | 95.130 | Sarcophilus_harrisii |
| ENSG00000184702 | SEPT5 | 100 | 91.111 | ENSSFOG00015014043 | sept5 | 98 | 90.258 | Scleropages_formosus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSSMAG00000006148 | sept5a | 99 | 85.185 | Scophthalmus_maximus |
| ENSG00000184702 | SEPT5 | 100 | 87.778 | ENSSMAG00000009333 | sept5b | 98 | 83.056 | Scophthalmus_maximus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSSDUG00000002398 | sept5a | 97 | 89.177 | Seriola_dumerili |
| ENSG00000184702 | SEPT5 | 100 | 86.667 | ENSSDUG00000006594 | sept5b | 99 | 81.843 | Seriola_dumerili |
| ENSG00000184702 | SEPT5 | 100 | 87.222 | ENSSLDG00000005541 | sept5b | 76 | 84.211 | Seriola_lalandi_dorsalis |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSSLDG00000003964 | sept5a | 97 | 87.446 | Seriola_lalandi_dorsalis |
| ENSG00000184702 | SEPT5 | 100 | 95.017 | ENSSPUG00000015346 | - | 100 | 94.587 | Sphenodon_punctatus |
| ENSG00000184702 | SEPT5 | 100 | 91.667 | ENSSPAG00000015627 | sept5a | 99 | 84.921 | Stegastes_partitus |
| ENSG00000184702 | SEPT5 | 100 | 88.333 | ENSSPAG00000000073 | sept5b | 93 | 86.246 | Stegastes_partitus |
| ENSG00000184702 | SEPT5 | 100 | 99.003 | ENSSSCG00000010128 | - | 100 | 92.118 | Sus_scrofa |
| ENSG00000184702 | SEPT5 | 100 | 95.556 | ENSTGUG00000008412 | - | 96 | 92.147 | Taeniopygia_guttata |
| ENSG00000184702 | SEPT5 | 100 | 85.000 | ENSTRUG00000018608 | sept5b | 98 | 80.165 | Takifugu_rubripes |
| ENSG00000184702 | SEPT5 | 100 | 90.556 | ENSTRUG00000016478 | sept5a | 97 | 88.728 | Takifugu_rubripes |
| ENSG00000184702 | SEPT5 | 100 | 84.444 | ENSTNIG00000013578 | sept5b | 99 | 79.730 | Tetraodon_nigroviridis |
| ENSG00000184702 | SEPT5 | 100 | 86.188 | ENSTNIG00000015926 | sept5a | 99 | 80.161 | Tetraodon_nigroviridis |
| ENSG00000184702 | SEPT5 | 100 | 99.336 | ENSUAMG00000011220 | - | 100 | 99.206 | Ursus_americanus |
| ENSG00000184702 | SEPT5 | 100 | 99.336 | ENSVVUG00000025380 | - | 88 | 99.148 | Vulpes_vulpes |
| ENSG00000184702 | SEPT5 | 100 | 95.017 | ENSXETG00000030888 | sept5 | 100 | 92.141 | Xenopus_tropicalis |
| ENSG00000184702 | SEPT5 | 100 | 91.111 | ENSXCOG00000002175 | sept5a | 99 | 89.112 | Xiphophorus_couchianus |
| ENSG00000184702 | SEPT5 | 100 | 85.083 | ENSXCOG00000016246 | sept5b | 97 | 82.759 | Xiphophorus_couchianus |
| ENSG00000184702 | SEPT5 | 100 | 85.083 | ENSXMAG00000016115 | sept5b | 97 | 82.759 | Xiphophorus_maculatus |
| ENSG00000184702 | SEPT5 | 100 | 91.111 | ENSXMAG00000014644 | sept5a | 98 | 89.112 | Xiphophorus_maculatus |
| Go ID | Go_term | PubmedID | Evidence | Category |
|---|---|---|---|---|
| GO:0003924 | GTPase activity | 21873635. | IBA | Function |
| GO:0003924 | GTPase activity | 21563316. | TAS | Function |
| GO:0005198 | structural molecule activity | 9611266. | TAS | Function |
| GO:0005515 | protein binding | 10321247.16767699.18330356.25416956. | IPI | Function |
| GO:0005525 | GTP binding | - | IEA | Function |
| GO:0005886 | plasma membrane | 10321247. | IDA | Component |
| GO:0005940 | septin ring | 21873635. | IBA | Component |
| GO:0008021 | synaptic vesicle | 21873635. | IBA | Component |
| GO:0008021 | synaptic vesicle | 10321247. | IDA | Component |
| GO:0015630 | microtubule cytoskeleton | 21873635. | IBA | Component |
| GO:0016080 | synaptic vesicle targeting | 10321247. | TAS | Process |
| GO:0017157 | regulation of exocytosis | 21873635. | IBA | Process |
| GO:0017157 | regulation of exocytosis | 10321247. | IMP | Process |
| GO:0030534 | adult behavior | - | ISS | Process |
| GO:0031105 | septin complex | 21873635. | IBA | Component |
| GO:0031105 | septin complex | - | ISS | Component |
| GO:0035176 | social behavior | - | ISS | Process |
| GO:0060090 | molecular adaptor activity | 21873635. | IBA | Function |
| GO:0061640 | cytoskeleton-dependent cytokinesis | 21873635. | IBA | Process |
| GO:2000300 | regulation of synaptic vesicle exocytosis | 21563316. | TAS | Process |