
| Protein ID | Domain | Pfam ID | E-value | Domain number | Total number |
|---|---|---|---|---|---|
| ENSP00000216123 | APOBEC_N | PF08210.11 | 4.4e-35 | 1 | 1 |
| ENSP00000411754 | APOBEC_N | PF08210.11 | 4.5e-35 | 1 | 1 |
| ENSP00000385741 | APOBEC_N | PF08210.11 | 6.2e-35 | 1 | 1 |
| ENSP00000483689 | APOBEC_N | PF08210.11 | 6.2e-35 | 1 | 1 |
| ENSP00000393520 | APOBEC_N | PF08210.11 | 3.8e-27 | 1 | 1 |
| ENSP00000482682 | APOBEC_N | PF08210.11 | 3.8e-27 | 1 | 1 |
| ENSP00000216123 | APOBEC_C | PF05240.14 | 1.5e-24 | 1 | 1 |
| ENSP00000411754 | APOBEC_C | PF05240.14 | 1.5e-24 | 1 | 1 |
| ENSP00000385741 | APOBEC_C | PF05240.14 | 1.9e-24 | 1 | 1 |
| ENSP00000483689 | APOBEC_C | PF05240.14 | 1.9e-24 | 1 | 1 |
| ENSP00000393520 | APOBEC_C | PF05240.14 | 1.1e-09 | 1 | 1 |
| ENSP00000482682 | APOBEC_C | PF05240.14 | 1.1e-09 | 1 | 1 |
| Transcript ID | Name | Length | RefSeq ID | Protein ID | Length | RefSeq ID | UniportKB ID |
|---|---|---|---|---|---|---|---|
| ENST00000613996 | APOBEC3H-207 | 465 | - | ENSP00000482682 | 154 (aa) | - | A0A087WZI3 |
| ENST00000401756 | APOBEC3H-202 | 1023 | XM_011529990 | ENSP00000385741 | 200 (aa) | XP_011528292 | Q6NTF7 |
| ENST00000421988 | APOBEC3H-203 | 592 | - | ENSP00000393520 | 154 (aa) | - | Q6NTF7 |
| ENST00000474235 | APOBEC3H-205 | 338 | - | - | - (aa) | - | - |
| ENST00000613677 | APOBEC3H-206 | 730 | - | ENSP00000483689 | 200 (aa) | - | UPI000188E67A |
| ENST00000348946 | APOBEC3H-201 | 1046 | - | ENSP00000216123 | 182 (aa) | - | Q6NTF7 |
| ENST00000442487 | APOBEC3H-204 | 988 | - | ENSP00000411754 | 183 (aa) | - | Q6NTF7 |
| Pathway ID | Pathway Name | Source |
|---|---|---|
| hsa05170 | Human immunodeficiency virus 1 infection | KEGG |
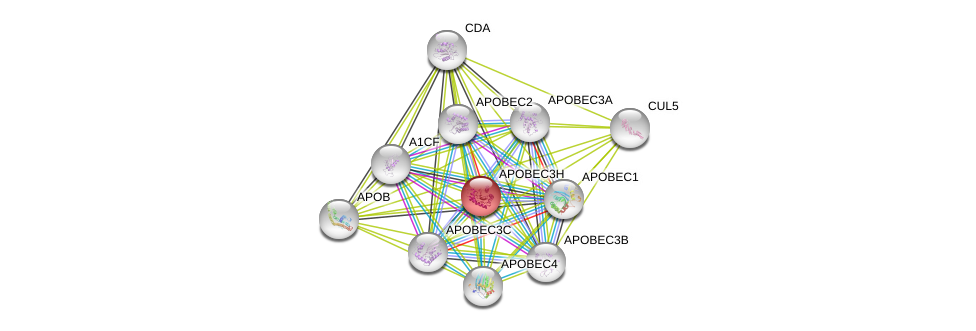
| Ensembl ID | Gene Symbol | Coverage | Identiy | Paralog | Gene Symbol | Coverage | Identiy |
|---|---|---|---|---|---|---|---|
| ENSG00000100298 | APOBEC3H | 99 | 33.163 | ENSG00000179750 | APOBEC3B | 90 | 32.911 |
| ENSG00000100298 | APOBEC3H | 99 | 34.759 | ENSG00000128394 | APOBEC3F | 88 | 36.025 |
| ENSG00000100298 | APOBEC3H | 74 | 41.045 | ENSG00000244509 | APOBEC3C | 71 | 40.288 |
| ENSG00000100298 | APOBEC3H | 97 | 33.862 | ENSG00000239713 | APOBEC3G | 92 | 33.161 |
| Ensembl ID | Gene Symbol | Coverage | Identiy | Ortholog | Gene Symbol | Coverage | Identiy | Species |
|---|---|---|---|---|---|---|---|---|
| ENSG00000100298 | APOBEC3H | 100 | 46.486 | ENSAMEG00000010818 | - | 76 | 46.486 | Ailuropoda_melanoleuca |
| ENSG00000100298 | APOBEC3H | 100 | 72.581 | ENSANAG00000025410 | APOBEC3H | 83 | 70.811 | Aotus_nancymaae |
| ENSG00000100298 | APOBEC3H | 99 | 48.889 | ENSBTAG00000007755 | APOBEC3H | 92 | 48.889 | Bos_taurus |
| ENSG00000100298 | APOBEC3H | 100 | 72.078 | ENSCJAG00000012524 | APOBEC3H | 100 | 69.481 | Callithrix_jacchus |
| ENSG00000100298 | APOBEC3H | 99 | 46.703 | ENSCAFG00000001361 | APOBEC3Z3 | 93 | 45.604 | Canis_familiaris |
| ENSG00000100298 | APOBEC3H | 99 | 46.154 | ENSCAFG00020002584 | - | 96 | 45.026 | Canis_lupus_dingo |
| ENSG00000100298 | APOBEC3H | 99 | 50.000 | ENSCHIG00000026580 | - | 95 | 50.000 | Capra_hircus |
| ENSG00000100298 | APOBEC3H | 98 | 51.124 | ENSTSYG00000032894 | - | 100 | 51.124 | Carlito_syrichta |
| ENSG00000100298 | APOBEC3H | 100 | 52.486 | ENSCPOG00000009257 | - | 86 | 52.809 | Cavia_porcellus |
| ENSG00000100298 | APOBEC3H | 100 | 74.522 | ENSCCAG00000008525 | APOBEC3H | 100 | 73.885 | Cebus_capucinus |
| ENSG00000100298 | APOBEC3H | 100 | 82.500 | ENSCATG00000030302 | APOBEC3H | 100 | 82.500 | Cercocebus_atys |
| ENSG00000100298 | APOBEC3H | 98 | 53.333 | ENSCLAG00000016615 | - | 88 | 53.333 | Chinchilla_lanigera |
| ENSG00000100298 | APOBEC3H | 100 | 80.874 | ENSCSAG00000006371 | APOBEC3H | 87 | 80.874 | Chlorocebus_sabaeus |
| ENSG00000100298 | APOBEC3H | 100 | 49.462 | ENSCGRG00001022976 | Apobec3 | 84 | 49.462 | Cricetulus_griseus_chok1gshd |
| ENSG00000100298 | APOBEC3H | 100 | 49.462 | ENSCGRG00000005621 | Apobec3 | 84 | 49.462 | Cricetulus_griseus_crigri |
| ENSG00000100298 | APOBEC3H | 99 | 50.811 | ENSDORG00000027271 | - | 99 | 42.857 | Dipodomys_ordii |
| ENSG00000100298 | APOBEC3H | 98 | 52.809 | ENSECAG00000007726 | APOBEC3H | 75 | 52.809 | Equus_caballus |
| ENSG00000100298 | APOBEC3H | 99 | 50.556 | ENSFCAG00000005251 | APOBEC3Z3 | 93 | 50.273 | Felis_catus |
| ENSG00000100298 | APOBEC3H | 99 | 51.111 | ENSSTOG00000007125 | - | 85 | 51.111 | Ictidomys_tridecemlineatus |
| ENSG00000100298 | APOBEC3H | 97 | 48.529 | ENSJJAG00000016840 | - | 73 | 48.529 | Jaculus_jaculus |
| ENSG00000100298 | APOBEC3H | 100 | 51.124 | ENSLAFG00000027404 | - | 95 | 51.124 | Loxodonta_africana |
| ENSG00000100298 | APOBEC3H | 100 | 83.766 | ENSMFAG00000031827 | APOBEC3H | 100 | 83.766 | Macaca_fascicularis |
| ENSG00000100298 | APOBEC3H | 100 | 84.416 | ENSMMUG00000001016 | APOBEC3H | 100 | 84.416 | Macaca_mulatta |
| ENSG00000100298 | APOBEC3H | 100 | 83.766 | ENSMNEG00000039259 | APOBEC3H | 100 | 83.766 | Macaca_nemestrina |
| ENSG00000100298 | APOBEC3H | 100 | 84.416 | ENSMLEG00000033417 | APOBEC3H | 100 | 84.416 | Mandrillus_leucophaeus |
| ENSG00000100298 | APOBEC3H | 100 | 48.128 | ENSMAUG00000011683 | Apobec3 | 72 | 47.568 | Mesocricetus_auratus |
| ENSG00000100298 | APOBEC3H | 99 | 48.901 | ENSMOCG00000021576 | Apobec3 | 83 | 48.333 | Microtus_ochrogaster |
| ENSG00000100298 | APOBEC3H | 100 | 48.108 | MGP_CAROLIEiJ_G0020029 | Apobec3 | 92 | 47.568 | Mus_caroli |
| ENSG00000100298 | APOBEC3H | 99 | 47.590 | ENSMUSG00000009585 | Apobec3 | 99 | 43.434 | Mus_musculus |
| ENSG00000100298 | APOBEC3H | 100 | 48.936 | MGP_PahariEiJ_G0020031 | - | 86 | 47.340 | Mus_pahari |
| ENSG00000100298 | APOBEC3H | 100 | 47.027 | MGP_SPRETEiJ_G0020927 | Apobec3 | 98 | 38.095 | Mus_spretus |
| ENSG00000100298 | APOBEC3H | 99 | 48.333 | ENSMPUG00000013239 | - | 78 | 48.333 | Mustela_putorius_furo |
| ENSG00000100298 | APOBEC3H | 100 | 48.333 | ENSNGAG00000018998 | Apobec3 | 93 | 48.333 | Nannospalax_galili |
| ENSG00000100298 | APOBEC3H | 100 | 87.978 | ENSNLEG00000015218 | - | 87 | 87.978 | Nomascus_leucogenys |
| ENSG00000100298 | APOBEC3H | 98 | 40.223 | ENSOCUG00000023913 | - | 81 | 40.223 | Oryctolagus_cuniculus |
| ENSG00000100298 | APOBEC3H | 99 | 49.444 | ENSOARG00000016290 | APOBEC3F | 96 | 49.444 | Ovis_aries |
| ENSG00000100298 | APOBEC3H | 100 | 95.455 | ENSPPAG00000036522 | APOBEC3H | 100 | 95.455 | Pan_paniscus |
| ENSG00000100298 | APOBEC3H | 99 | 51.381 | ENSPPRG00000003941 | - | 93 | 50.543 | Panthera_pardus |
| ENSG00000100298 | APOBEC3H | 92 | 35.260 | ENSPPRG00000003675 | - | 88 | 34.682 | Panthera_pardus |
| ENSG00000100298 | APOBEC3H | 99 | 51.381 | ENSPTIG00000010283 | - | 93 | 50.543 | Panthera_tigris_altaica |
| ENSG00000100298 | APOBEC3H | 100 | 95.455 | ENSPTRG00000014390 | APOBEC3H | 100 | 95.455 | Pan_troglodytes |
| ENSG00000100298 | APOBEC3H | 100 | 84.066 | ENSPANG00000010774 | APOBEC3H | 99 | 83.660 | Papio_anubis |
| ENSG00000100298 | APOBEC3H | 100 | 45.161 | ENSPEMG00000000456 | Apobec3 | 76 | 44.624 | Peromyscus_maniculatus_bairdii |
| ENSG00000100298 | APOBEC3H | 100 | 88.525 | ENSPPYG00000011834 | - | 87 | 88.525 | Pongo_abelii |
| ENSG00000100298 | APOBEC3H | 98 | 42.135 | ENSPCAG00000015655 | - | 99 | 41.111 | Procavia_capensis |
| ENSG00000100298 | APOBEC3H | 70 | 37.008 | ENSPVAG00000024267 | - | 93 | 37.008 | Pteropus_vampyrus |
| ENSG00000100298 | APOBEC3H | 98 | 51.124 | ENSPVAG00000009462 | - | 98 | 51.124 | Pteropus_vampyrus |
| ENSG00000100298 | APOBEC3H | 70 | 37.008 | ENSPVAG00000024161 | - | 93 | 37.008 | Pteropus_vampyrus |
| ENSG00000100298 | APOBEC3H | 100 | 48.663 | ENSRNOG00000016852 | Apobec3 | 87 | 48.663 | Rattus_norvegicus |
| ENSG00000100298 | APOBEC3H | 100 | 81.967 | ENSRBIG00000029088 | APOBEC3H | 100 | 81.818 | Rhinopithecus_bieti |
| ENSG00000100298 | APOBEC3H | 100 | 81.421 | ENSRROG00000024699 | APOBEC3H | 100 | 81.000 | Rhinopithecus_roxellana |
| ENSG00000100298 | APOBEC3H | 99 | 70.492 | ENSSBOG00000030005 | APOBEC3H | 100 | 69.504 | Saimiri_boliviensis_boliviensis |
| ENSG00000100298 | APOBEC3H | 99 | 50.556 | ENSSSCG00000000091 | APOBEC3B | 83 | 50.556 | Sus_scrofa |
| ENSG00000100298 | APOBEC3H | 95 | 50.000 | ENSTBEG00000001532 | - | 98 | 50.000 | Tupaia_belangeri |
| ENSG00000100298 | APOBEC3H | 100 | 51.099 | ENSTTRG00000001266 | - | 100 | 51.099 | Tursiops_truncatus |
| ENSG00000100298 | APOBEC3H | 99 | 47.872 | ENSUAMG00000004427 | - | 83 | 47.872 | Ursus_americanus |
| ENSG00000100298 | APOBEC3H | 99 | 50.829 | ENSUMAG00000014435 | - | 91 | 50.829 | Ursus_maritimus |
| ENSG00000100298 | APOBEC3H | 71 | 51.538 | ENSVPAG00000011108 | - | 98 | 51.538 | Vicugna_pacos |
| Go ID | Go_term | PubmedID | Evidence | Category |
|---|---|---|---|---|
| GO:0000932 | P-body | 22915799. | IDA | Component |
| GO:0003723 | RNA binding | 21873635. | IBA | Function |
| GO:0004126 | cytidine deaminase activity | 21873635. | IBA | Function |
| GO:0004126 | cytidine deaminase activity | 16571802. | IDA | Function |
| GO:0005515 | protein binding | 22915799. | IPI | Function |
| GO:0005634 | nucleus | 21873635. | IBA | Component |
| GO:0005634 | nucleus | 18779051. | IDA | Component |
| GO:0005737 | cytoplasm | 21873635. | IBA | Component |
| GO:0005737 | cytoplasm | 18779051.21835787. | IDA | Component |
| GO:0008270 | zinc ion binding | - | IEA | Function |
| GO:0009972 | cytidine deamination | - | IEA | Process |
| GO:0010529 | negative regulation of transposition | 18779051.20062055. | IDA | Process |
| GO:0010529 | negative regulation of transposition | 18827027. | IMP | Process |
| GO:0016554 | cytidine to uridine editing | 21873635. | IBA | Process |
| GO:0045087 | innate immune response | - | IEA | Process |
| GO:0045869 | negative regulation of single stranded viral RNA replication via double stranded DNA intermediate | 18779051.21835787. | IDA | Process |
| GO:0048525 | negative regulation of viral process | 18827027. | IMP | Process |
| GO:0051607 | defense response to virus | 21835787.22457529.22915799. | IDA | Process |
| GO:0070383 | DNA cytosine deamination | 16571802.21835787. | IDA | Process |
| GO:0080111 | DNA demethylation | 21873635. | IBA | Process |
| Cancer | Chr | Position | Mutation Type | dbSNP | Protein-change | Allele Freq | RBD |
|---|---|---|---|---|---|---|---|
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| BLCA | |||||||
| CESC | |||||||
| CESC | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| COAD | |||||||
| ESCA | |||||||
| GBM | |||||||
| GBM | |||||||
| GBM | |||||||
| HNSC | |||||||
| HNSC | |||||||
| LGG | |||||||
| LGG | |||||||
| LIHC | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUAD | |||||||
| LUSC | |||||||
| LUSC | |||||||
| PAAD | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| SKCM | |||||||
| STAD | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC | |||||||
| UCEC |
| Cancer | Type | Freq | Q-value |
|---|---|---|---|
| PCPG | |||
| THCA | |||
| THYM |
| Cancer | P-value | Q-value |
|---|---|---|
| THYM | ||
| KIRC | ||
| SARC | ||
| ACC | ||
| HNSC | ||
| SKCM | ||
| BRCA | ||
| BLCA | ||
| CESC | ||
| LAML | ||
| UCEC | ||
| LIHC | ||
| LGG | ||
| CHOL | ||
| THCA | ||
| LUAD | ||
| UVM | ||
| OV |