
| PID | Title | Article | Time | Author | Doi |
|---|---|---|---|---|---|
| 19587378 |
| Transcript ID | Name | Length | RefSeq ID | Protein ID | Length | RefSeq ID | UniportKB ID |
|---|---|---|---|---|---|---|---|
| ENST00000543772 | PSMC1-202 | 1389 | XM_017021470 | ENSP00000445147 | 367 (aa) | XP_016876959 | P62191 |
| ENST00000261303 | PSMC1-201 | 4448 | - | ENSP00000261303 | 440 (aa) | - | P62191 |
| ENST00000557357 | PSMC1-207 | 581 | - | - | - (aa) | - | - |
| ENST00000553835 | PSMC1-203 | 390 | - | ENSP00000452049 | 84 (aa) | - | G3V4X1 |
| ENST00000554624 | PSMC1-204 | 577 | - | - | - (aa) | - | - |
| ENST00000555787 | PSMC1-206 | 2246 | - | - | - (aa) | - | - |
| ENST00000555679 | PSMC1-205 | 606 | - | - | - (aa) | - | - |
| Ensembl ID | Gene Symbol | Coverage | Identiy | Ortholog | Gene Symbol | Coverage | Identiy | Species |
|---|---|---|---|---|---|---|---|---|
| ENSG00000100764 | PSMC1 | 99 | 90.361 | WBGene00004502 | rpt-2 | 94 | 85.337 | Caenorhabditis_elegans |
| ENSG00000100764 | PSMC1 | 98 | 54.878 | WBGene00004503 | rpt-3 | 89 | 52.162 | Caenorhabditis_elegans |
| ENSG00000100764 | PSMC1 | 95 | 62.500 | WBGene00004501 | rpt-1 | 78 | 47.198 | Caenorhabditis_elegans |
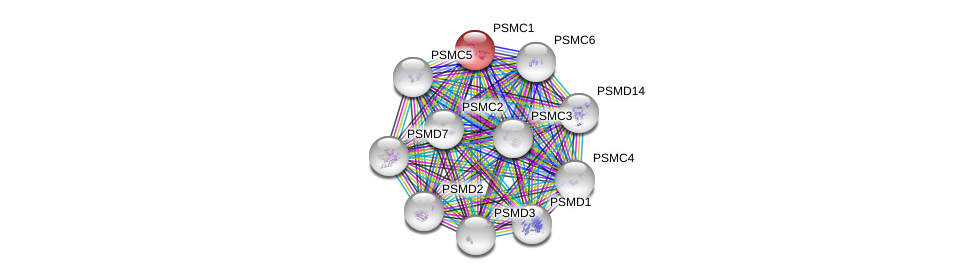
| ENSG00000100764 | PSMC1 | 99 | 65.060 | ENSG00000087191 | PSMC5 | 83 | 55.932 | Homo_sapiens |
| ENSG00000100764 | PSMC1 | 95 | 61.250 | ENSG00000165916 | PSMC3 | 90 | 45.119 | Homo_sapiens |
| ENSG00000100764 | PSMC1 | 95 | 60.000 | ENSG00000161057 | PSMC2 | 84 | 47.321 | Homo_sapiens |
| ENSG00000100764 | PSMC1 | 98 | 56.098 | ENSG00000100519 | PSMC6 | 99 | 46.970 | Homo_sapiens |
| ENSG00000100764 | PSMC1 | 99 | 84.337 | YDL007W | RPT2 | 95 | 73.317 | Saccharomyces_cerevisiae |
| ENSG00000100764 | PSMC1 | 98 | 60.976 | YDR394W | RPT3 | 87 | 49.330 | Saccharomyces_cerevisiae |
| ENSG00000100764 | PSMC1 | 99 | 62.651 | YGL048C | RPT6 | 80 | 51.235 | Saccharomyces_cerevisiae |
| ENSG00000100764 | PSMC1 | 95 | 61.250 | YKL145W | RPT1 | 58 | 53.532 | Saccharomyces_cerevisiae |
| Go ID | Go_term | PubmedID | Evidence | Category |
|---|---|---|---|---|
| GO:0000165 | MAPK cascade | - | TAS | Process |
| GO:0000209 | protein polyubiquitination | - | TAS | Process |
| GO:0000502 | proteasome complex | 17323924. | IDA | Component |
| GO:0002223 | stimulatory C-type lectin receptor signaling pathway | - | TAS | Process |
| GO:0002479 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent | - | TAS | Process |
| GO:0003723 | RNA binding | 22658674. | HDA | Function |
| GO:0005515 | protein binding | 9452483.16189514.16990800.19060904.19490896.22901813.23455924.25416956.26496610. | IPI | Function |
| GO:0005524 | ATP binding | - | IEA | Function |
| GO:0005634 | nucleus | 21630459. | HDA | Component |
| GO:0005654 | nucleoplasm | - | IDA | Component |
| GO:0005654 | nucleoplasm | - | TAS | Component |
| GO:0005829 | cytosol | - | IDA | Component |
| GO:0005829 | cytosol | - | TAS | Component |
| GO:0006457 | protein folding | - | TAS | Process |
| GO:0006521 | regulation of cellular amino acid metabolic process | - | TAS | Process |
| GO:0008540 | proteasome regulatory particle, base subcomplex | 21873635. | IBA | Component |
| GO:0010972 | negative regulation of G2/M transition of mitotic cell cycle | - | TAS | Process |
| GO:0016020 | membrane | 19946888. | HDA | Component |
| GO:0016579 | protein deubiquitination | - | TAS | Process |
| GO:0016887 | ATPase activity | 1429620. | ISS | Function |
| GO:0017025 | TBP-class protein binding | 21873635. | IBA | Function |
| GO:0022624 | proteasome accessory complex | - | ISS | Component |
| GO:0031145 | anaphase-promoting complex-dependent catabolic process | - | TAS | Process |
| GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process | - | TAS | Process |
| GO:0033209 | tumor necrosis factor-mediated signaling pathway | - | TAS | Process |
| GO:0036402 | proteasome-activating ATPase activity | 21873635. | IBA | Function |
| GO:0038061 | NIK/NF-kappaB signaling | - | TAS | Process |
| GO:0038095 | Fc-epsilon receptor signaling pathway | - | TAS | Process |
| GO:0043161 | proteasome-mediated ubiquitin-dependent protein catabolic process | - | TAS | Process |
| GO:0043488 | regulation of mRNA stability | - | TAS | Process |
| GO:0043687 | post-translational protein modification | - | TAS | Process |
| GO:0050852 | T cell receptor signaling pathway | - | TAS | Process |
| GO:0055085 | transmembrane transport | - | TAS | Process |
| GO:0060071 | Wnt signaling pathway, planar cell polarity pathway | - | TAS | Process |
| GO:0061418 | regulation of transcription from RNA polymerase II promoter in response to hypoxia | - | TAS | Process |
| GO:0070498 | interleukin-1-mediated signaling pathway | - | TAS | Process |
| GO:0090090 | negative regulation of canonical Wnt signaling pathway | - | TAS | Process |
| GO:0090263 | positive regulation of canonical Wnt signaling pathway | - | TAS | Process |
| GO:1901215 | negative regulation of neuron death | - | IEA | Process |
| GO:1901800 | positive regulation of proteasomal protein catabolic process | - | IEA | Process |
| GO:1901990 | regulation of mitotic cell cycle phase transition | - | TAS | Process |
| GO:1902036 | regulation of hematopoietic stem cell differentiation | - | TAS | Process |